1. Introduction
This manual is written primarily for national GLASS data managers. It can also be used by data managers at laboratories serving surveillance sites participating in GLASS. It describes how to use WHONET in handling data for GLASS. WHONET supports output of data to GLASS exchange format for uploading to the GLASS IT platform. This is not the complete WHONET manual describing all its functionalities and features. The complete WHONET manual (including BacLink tutorials) can be downloaded from the WHONET home page at www.whonet.org.
WHONET supports participation in GLASS as:
§ it is freeware, downloadable from the WHONET Home Page, www.whonet.org; and
§ it supports output to the GLASS data exchange format.
WHONET includes Data Check Feedback Reports, e.g.
§ it gives direct feedback on microbiological aspects for the isolate level (e. g. alert on Methicillin-resistant Staphylococcus aureus; Penicillin non-susceptible Streptococcus pneumoniae to allow possible interventions on site; and
§ it checks GLASS aggregated data validity and completeness, producing feedback reports indicating if it is necessary to correct data or make them complete before uploading to the GLASS IT platform.
Section 2: “Installing WHONET” is intended for national and local users of the WHONET software.
Section 3: “Laboratory configuration” and Section 4: “WHONET data entry for GLASS” are intended for users who will enter data into WHONET. Data entry can be accomplished at the national level by technical staff of the network coordinating institution who receives data submitted on paper forms from participating facilities. In the case of data entry into WHONET at the level of participating facilities, electronic WHONET data files are sent to the national level; at the national level WHONET data files from different facilities are combined and aggregated to the GLASS exchange format.
Note: If participating facilities already have laboratory information systems or susceptibility test instruments with the desired surveillance data, it is usually possible to avoid manual data entry through the use of BacLink – WHONET’s data import module – to capture and standardize data from existing information resources. Further information about BacLink is provided as an Annex in this manual and can also be found in the BacLink tutorial BacLink 1 – Getting started which is available from http://www.whonet.org/documentation.html.
Sections 5 through 8 are intended for use by national surveillance system coordinators responsible for collation, analysis, validation and submission of surveillance data to the GLASS IT platform.
For comments and questions on WHONET use, please write to help@whonet.org.
2. Installing WHONET
The software is available from the WHONET Home Page at www.whonet.org. Click on “Software” and download WHONET 2016 (Desktop version).
WHONET requires a computer running Microsoft Windows versions XP or later, and also requires that Windows .NET version 4.0 is installed. After downloading the software, double-click on the downloaded file and follow the instructions on the screen (“Run”, “OK”, etc.). If you receive a message about “insufficient administrative rights” to install the software, you will need to contact your facility’s system administrator to install the software for you or to grant you the necessary local administration rights temporarily.
The default location for the WHONET installation is C:\WHONET\. You may change this as you wish. For example, if you would like to use WHONET on a network drive to permit shared use by multiple staff members, you can install the software to a common drive such as T:\WHONET\.
In case you already have an older version of WHONET (prior to “WHONET 2016”) on your PC, then you may install the updated WHONET applications side by side in different folders and copy your data and configuration files to the new folders. This method will retain your ability to use the older version of the software. Alternatively, you can use the browse feature to locate the configuration files without copying them.
After installing the programme, you will find the WHONET and BacLink icons on your computer’s desktop.
Note: If you have an antivirus, or system protection software installed such as Microsoft SmartScreen, Norton Antivirus, Symantec Antivirus, etc., it is possible that you may encounter a warning message similar to the following:
![]()
This message appears because the software is new and has not been installed on many systems, so the antivirus software notifies you that you should take extra precaution. It does not indicate a problem with the installation package. To proceed, you must follow your system’s prompts to continue with the installation. It may be necessary to open your computer’s downloads folder in order to locate the installation package.
Additionally, you may wish to scan the file using your antivirus software before installing the application. You should also verify that the installer is from the verified publisher “Brigham and Women’s Hospital.” This last point ensures that the installation package is authentic and has not been modified since it was created.
3. Laboratory configuration
3.1 Creating a new laboratory configuration
National data managers who receive paper forms from participating surveillance sites need to enter data at the national level. The first step is to create a GLASS data format-compatible laboratory. You may wish to create one laboratory for each participating surveillance site (see section 3.2 for “Copying existing laboratory configuration”) or a single “national” laboratory that can be utilized for analysing and managing data from any of the participating facilities in the country.
Where participating surveillance sites submit electronic WHONET data files to the national level, the data manager may first want to create a GLASS laboratory to combine data into one WHONET file (line-level data) and subsequently aggregate data to the GLASS exchange format (see section 6 for exporting WHONET files to the GLASS exchange format).
To configure a GLASS laboratory:
Ø Start WHONET, the screen below appears, click “Cancel”
Then the main WHONET screen appears.
Ø Click “File”, select “GLASS“ and click “New laboratory”, the following screen appears:
Ø Indicate your country, laboratory name and laboratory code, then click “OK”.
The screen below appears:
Ø Choose your configuration type. Data entered through either method will be usable for GLASS export. Indeed, WHONET files created by any method can be exported to GLASS formats. The minimal GLASS configuration reflects the GLASS standards outlined in the WHO GLASS Manual for Early Implementation.[1] The standard configuration will allow for more extensive data entry and analysis options in addition to GLASS requirements.
The screen below appears:
Ø Select the guidelines used at the laboratory and click “OK”.
With the minimal GLASS configuration WHONET will create a new laboratory with the recommended set of GLASS antibiotics, data fields and codes. You will then be prompted with the following screen.
Ø If you want to review the configuration and make any additional modifications, select “Yes” and follow instructions in section 3.3 and Annex 1. Otherwise, select “No” to proceed directly with WHONET data entry and analysis.
Before you start with data entry we recommend that you review to check if it is necessary to make changes to the antibiotic interpretative breakpoints. For details of how to make any additional changes please refer to section 3.3 and Annex 1.
3.2 Copying an existing laboratory configuration
If the national data manager is responsible for working with multiple facilities, the “Copy laboratory” feature can be used to clone the created GLASS laboratory to new ones, assigning a new name and code to each additional facility created. This is described in further detail below.
Ø Choose from main menu, “File”, “Open laboratory”.
Ø To copy an existing laboratory configuration e.g. SWE-005, select SWE-005 from the screen below, and click on “Copy laboratory”.
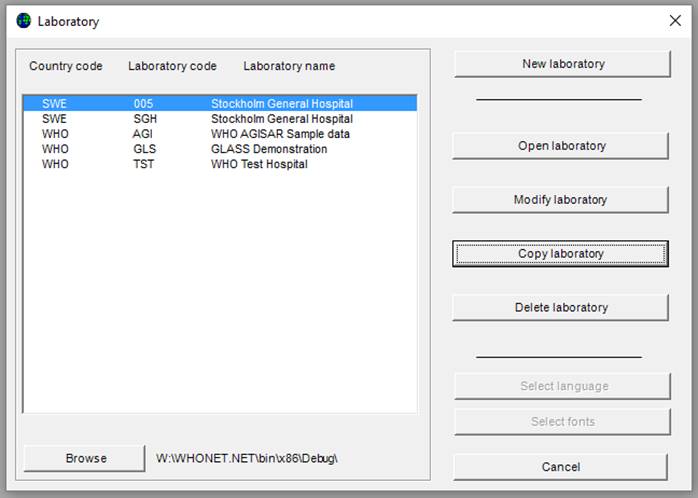
The screen below will appear:
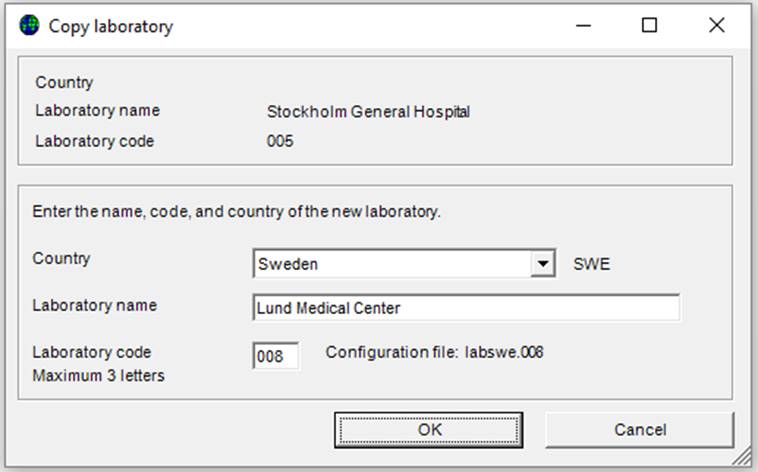
Ø Enter the country, laboratory name and laboratory code (numbers) and click “OK”.
3.3 Modifying laboratory configuration
When using the feature “Create a new GLASS laboratory”, WHONET defines a laboratory configuration with a list of antibiotics and data fields. However, the data manager may wish to further customize the configuration to more closely match data entry, analysis and reporting needs on site and at the national level.
Options include:
§ General information: The user may modify the laboratory code, full laboratory name and country code. The user can also specify whether isolates of human origin will be collected primarily or whether human, animal, food and/or environmental isolates are to be collected in one database.
§ Data fields: When the “Create a new GLASS laboratory” feature was used, WHONET created a list of “minimal GLASS” or “standard WHONET” data fields, such as “Identification number”, “Specimen type” and “Pathogen”. The user may add or remove data fields from this list to more closely match data management preferences, for example “Patient last name” or “Diagnosis”.
§ Locations: A list of patient care areas may be created for the laboratory configuration e.g. “Emergency Room”, “General medical ward” and “Surgical intensive care unit”. This will be particularly useful if the user plans to use WHONET for purposes of clinical reporting or data analysis on site. If data for this variable are to be combined at the national level with data from different facilities, a standardized list of patient care areas must be implemented by the coordinating centre.
§ Antibiotics: Antibiotics may be added or removed from the antibiotic list established for the laboratory configuration so that it more closely matches the set of antibiotics tested on site or considered under surveillance at the national level. “Antibiotic panels” may be defined, which control the organism-specific list of antibiotics displayed to the user during data entry or “antibiotic profiles” used in the WHONET data analysis module. The breakpoints automatically defined by WHONET can be reviewed and edited in this module.
Further details on these configuration features can be found in Annex 1.
4. WHONET data entry for GLASS
4.1 General
Before starting “Data entry” the user should open the corresponding laboratory configuration.
Ø Click on main menu, “File”, “Open laboratory”. Select laboratory and click on “Open laboratory”.
WHONET uses the data structure defined in the current laboratory configuration and will use the breakpoints as defined in the laboratory configuration.
Ø Select a new data file or open an existing file. To make a new file, select “Data Entry” from the menu bar and then select “New Data File”.
The screen Data entry (below) appears.
Ø Enter the name of the file to be created, for example “2015SWE.005” or “2015GLASS.008”
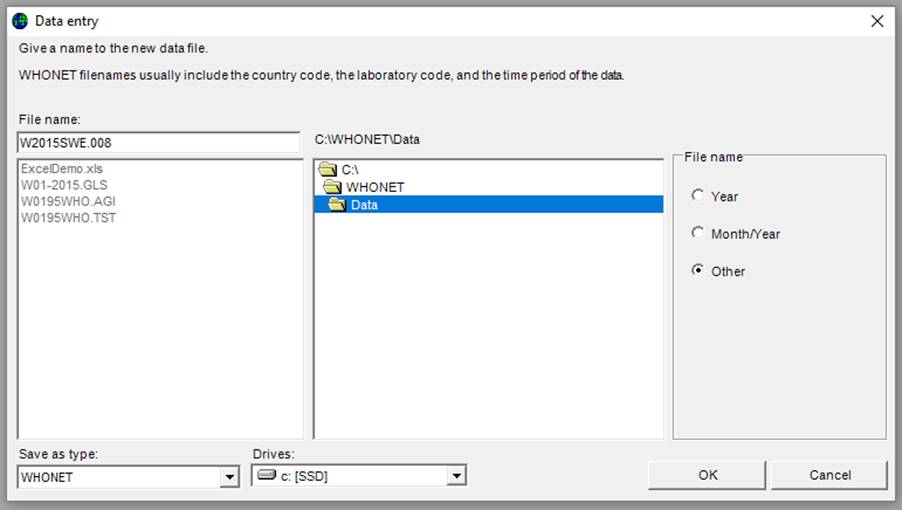
By default, data files are stored in the C:\WHONET\Data\ folder, but the user may use the “Drives” and “Folder” boxes to specify a preferred data file location.
Ø Click OK to proceed with data entry.
Once a data file has been opened or created, the “Data entry” screen (below) will appear.
Data are entered in the fields on the left half of the screen. The following data fields will appear if the “minimal GLASS configuration” was selected.
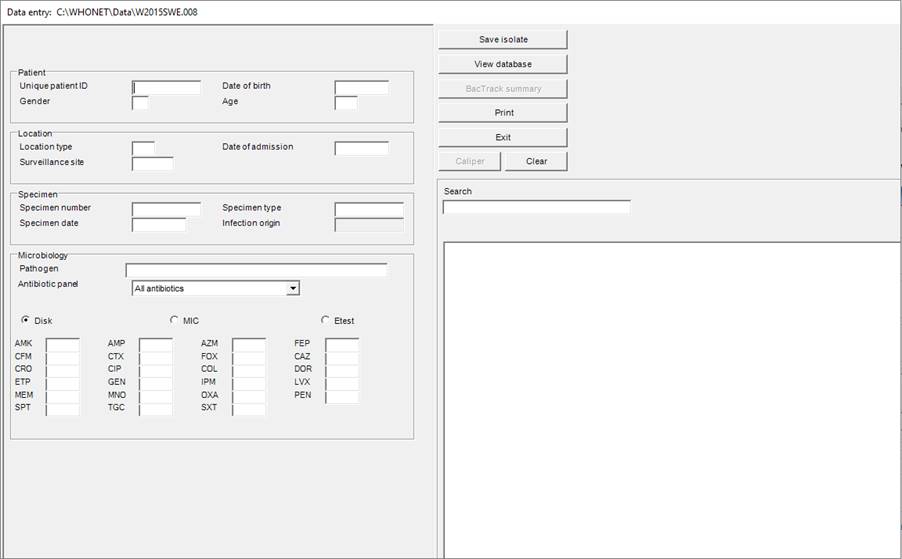
General explanation for data entry:
When the cursor jumps to a field for data entry, brief instructions and recommended data codes for that field appear at the lower right of the screen.
After entering data in one field, there are four ways of moving to the next field:
Ø press the <Enter> key; or
Ø press the <Tab> key; or
Ø press the arrow keys; or
Ø use the mouse.
Enter dates in the same format as the default format on the computer in use, namely as day/month/year or month/day/year or year/month/day. After entering a date and moving to the next field, the user should check that the date has been interpreted correctly – WHONET automatically converts the numeric month to the name of the month. When entering a date, the year can be entered as a 2- or 4-digit date. The numbers indicating day, month and year must be separated by a /, hyphen, or space.
4.2 Field-specific comments for GLASS minimal configuration
In the following section the variables of the GLASS minimal configuration laboratory are described. Most of them are required for generating the GLASS exchange format (GLASS aggregated data files). If the user has configured the laboratory to requirements different from the GLASS protocol, additional data entry fields may appear.
Note: GLASS is collecting data not only on “positive” results (GLASS priority pathogens) but also on the overall number of clinical specimens submitted to the laboratory for species isolation and identification. That means, for example for blood specimens, information on isolates other than the GLASS priority pathogens and also “negative” results should be included in the aggregated GLASS data submission. WHONET has implemented this approach in the GLASS sample file, providing the information is entered on site; the overall number of samples for the respective specimen is aggregated in the sample file. For more details on data entry in WHONET see below under “Pathogen”. More information on the approach and methodology can be found in the GLASS Manual for Early Implementation[2] and the GLASS guide to preparing aggregated AMR data files.[3]
Unique patient identifier
Enter a unique patient identifier (ID) for each patient with a maximum of 12 characters. Several specimens and isolates can be assigned to each patient and corresponding unique ID. The unique patient ID is required for de-duplication that is automatically done by WHONET software when generating the GLASS exchange format (see also section 6.3).
Date of birth
To enter the date of birth see instructions on the previous page regarding the format. Alternatively the user may wish to enter the patient’s age (see “Age”). When generating the GLASS exchange format, age will be aggregated into the GLASS pre-defined age group category.
Gender
Enter “m” for male and “f” for female. Missing values will be reported in the GLASS aggregated data file as unknown.
Age
The option of entering the patient’s age exists. However, if the patient’s date of birth has been entered, the patient’s age will automatically be calculated and inserted in the age field when the specimen date is entered. When generating the GLASS exchange format, age will be grouped in the GLASS pre-defined age group category.
Date of admission
Enter the date of hospital (inpatient) admission (see previous page for date format). Date of admission is a field required to calculate “hospital origin”.
Location type
Select the location type (ward type). It is important to indicate the patient’s location as the calculation of the GLASS variable “origin” (community or hospital, see “infection origin” below) is based on an algorithm in the WHONET software that requires the variable’s location type. As a minimum enter outpatient or inpatient respectively. For hospitalized patients, the “date of admission” and “specimen date” are also required. Missing values will result in unknown “origin” in the GLASS aggregated data file.
Infection origin
The GLASS protocol defines “origin” as community- or hospital-acquired infection (see also the GLASS Manual for Early Implementation and the GLASS document “ A guide to preparing aggregated AMR data files” for more details). WHONET software calculates “Infection origin” using a combination of data entry fields. In order for the software to calculate this field, three data fields are needed: location type; and for hospitalized patients - specimen date and the date of admission. The rules for calculating the “Infection origin” (corresponding to the GLASS “origin”) are as follows: All location types of “out” (coded value for outpatient) will result in the isolate being reported to GLASS as “Community”. If admission date and specimen date are provided, and location type is “in” (coded value for inpatient), the system will verify that the admission date occurred more than two days prior to the specimen’s collection. If this is true, the isolate will be marked “Hospital”, otherwise it will be marked “Community”. If the system cannot determine the “Infection origin”, usually due to missing data, it will be reported as “Unknown” to GLASS.
Surveillance site
Enter the surveillance site (maximum 3 letters laboratory code) or choose “other” in the event that the patient has not been cared for at the surveillance site.
Specimen number
Enter a number for each specimen with a maximum of 12 characters.
The specimen number may be helpful for data validation purposes when, for example, several isolates from the same patient are entered.
Specimen date
Enter the date the specimen was taken (see previous page for date format). This field is required to calculate the “Infection origin”.
Specimen type
Enter the specimen type from the GLASS list that is displayed on the right half of the screen. Two lists are available to the user – the short “GLASS list” with four specimen types (“Blood”, “Genital”, “Stool” and “Urine”); or the “Standard WHONET list” with a much larger number of specimen types e.g. “Cerebrospinal fluid”, “Sputum”, “Joint fluid”, etc. Choose the list that is most appropriate for your needs.
Pathogen
The 3-character WHONET organism code should be entered here, or selected from the GLASS pathogen list on the right half of the screen that is displayed by default for the minimal GLASS configuration. To view the standard or extended list and to enter information on isolates other than the GLASS priority pathogens, select ”standard” or “extended” respectively from the drop down option. To enter data for specimens submitted to the laboratory but with “negative” results (“no isolate growth”) leave the pathogen field blank or use the WHONET code “xxx” signifying “No growth”, and press the “Save isolate” button on the upper right screen and proceed as prompted. The specimen will be correctly included in the GLASS sample file. More information on the approach and methodology can be found in the GLASS Manual for Early Implementation and the GLASS document “A guide to preparing aggregated AMR data files”.
Antibiotic panel
The antibiotic panel displayed by default shows the antibiotics for the selected organism proposed by GLASS for reporting. If the user changes the panel to “All antibiotics”, then it is possible to select from any of the antibiotics on the list.
Susceptibility results and the list of antibiotics
To enter susceptibility results, first click on the appropriate test method – disk diffusion, MIC or ETestÒ. The list of antibiotics for that test method appears. Each time a result is entered press <Enter>: the cursor then jumps to the next antibiotic on the antibiotic panel for the organism tested.
Entering susceptibility results
WHONET allows the entry of quantitative results (e.g. 13 mm, 64 µg/ml) or the entry of qualitative results (R = resistant, I = intermediate, S = susceptible).
The lowest possible zone diameter is 6 mm. If 0 mm is entered (indicating no inhibition), WHONET will automatically change this to 6 mm.
For off-scale MIC values the following, for example, may be entered: <=.5, >64.
If MIC results from the test of a drug combination are being entered (e.g. trimethoprim/sulfamethoxazole), enter the result of the first or principal agent. These concentrations usually follow the 1, 2, 4, 8 … doubling-dilution series.
Saving the isolate information
When all the data for an isolate have been entered, click on “Save isolate” (or press Alt-S). The data will be saved and the “Data entry” screen will be cleared so that the data for the next isolate can be entered.
WHONET will ask whether the user wants to:
§ Save the isolate
§ Save and continue with the same isolate
§ Save and continue with the same patient
Click on one of these options to save or on “Cancel”, if the decision is not to save the record.
Exit data entry
When all data have been entered, click on “Exit” to return to the main WHONET screen.
4.3 View database
Ø From menu choose, “Data entry”, “Open data file”, select data file and click on “Open”.
Ø To check the records once data entry has been performed, click on “View database”: the screen below appears.
Ø To make changes, select the record to be changed and click “Edit isolate.” The data entry screen will appear again and changes can be entered. Click “Save isolate” to save these changes.
Note: The calculated variable “Infection origin” is only visible in the data entry screen and not in the view database.
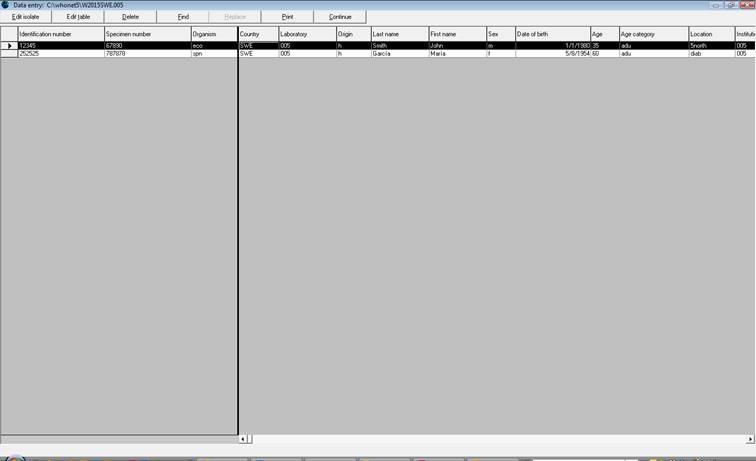
Ø To leave the screen above, click on “Continue” and “Exit”.
5. Data collection at the national level
The previous sections covered the entry of data into WHONET. This can be accomplished by staff at each of the surveillance sites participating in the national surveillance system or alternatively at the national level by the surveillance system coordinators utilizing paper forms or electronic records received from participating facilities.
If data entry is accomplished by the participating facilities, then data entry staff should periodically send their WHONET data files securely to the national programme coordinators. For purposes of GLASS data preparation, annual data submissions to the national coordinators may prove sufficient. However, to support the needs of the national surveillance programme, more frequent submissions, for example monthly, are recommended in order to recognize and respond to emerging threats.
If facilities already have electronic data stores, for example in a laboratory information system or a laboratory organism and identification susceptibility test instrument, then BacLink – WHONET’s data import module – can usually be utilized to import the data into WHONET. Further details are available in the BacLink tutorials, starting with BacLink 1 – Getting started which is available from http://www.whonet.org/documentation.html. BacLink can be run at the local level directly by facility staff members. Alternatively, national network coordinators can receive raw data files from participating facilities and convert these into WHONET files using BacLink at the national level, as introduced in Annex 2 and described further in a series of BacLink tutorials.
6. Exporting WHONET files to the GLASS exchange format
There are 2 scenarios for WHONET files:
§ Data have been entered manually at the national level following the instructions provided in the previous sections; or
§ electronic files in WHONET format have been received from participating surveillance sites.
Manual data entry
After manual data entry the files are saved in WHONET format. Before uploading them to the GLASS user interface, they must be exported to the GLASS data exchange format. It is possible that the user may have national WHONET data stored in a single large WHONET file, or alternatively, the user may have separate WHONET files representing various facilities, specimens and time periods. If multiple files exist, the user may first wish to combine them together into a single large WHONET file using the procedure described below (section 6.1).
Electronic data entry
After receiving electronic WHONET data files from participating surveillance sites the user may first want to combine data files from more than one surveillance site into a single large WHONET file. However, surveillance sites may use different WHONET software versions and different laboratory configurations. If the laboratories are testing exactly the same antibiotics with the same set of breakpoints, the data files can be combined using any one of the corresponding laboratory configurations. On the other hand, if the laboratories are testing different antibiotics or using different breakpoints, then a new “national” laboratory configuration should be created and customized that includes a list of all antibiotics tested (with the corresponding breakpoints), as mentioned earlier in Sections 3.1 and 3.3.
6.1 Combining WHONET files
We first describe how to combine WHONET files from different surveillance sites into one single large WHONET file before aggregation of data.
Ø To combine data files in a preferred configuration, select “Open laboratory” and choose the laboratory with the corresponding configuration.
Ø Click on “Data entry” from the main WHONET screen. Choose “Combine or export data files”. Indicate the data files that are to be combined and enter the name of the new data file being created. The default option “Save as type” is WHONET.
Ø Then click on “Combine”.
Ø When WHONET finishes, the original data files will be unchanged, but the user will now have a new large data file that includes all of the results from the original files. The new WHONET data file is stored per default in the C:\WHONET\Data folder.
6.2 Aggregation of WHONET files
The next steps describe the aggregation of one or several data files directly into the GLASS exchange format. The created data file will be stored per default in the C:\WHONET\Output folder.
Ø From the main WHONET menu, again select “Open laboratory” and choose the laboratory with the corresponding configuration created previously.
Ø From the main WHONET menu, select “Data entry”, “Combine or export data files”.
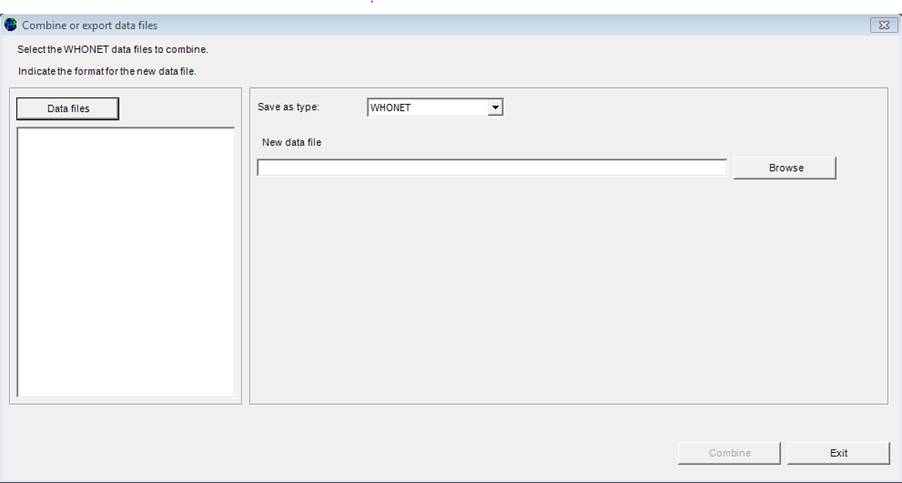
Ø Click on “Data files”.
The
next screen will appear (example file is displayed): 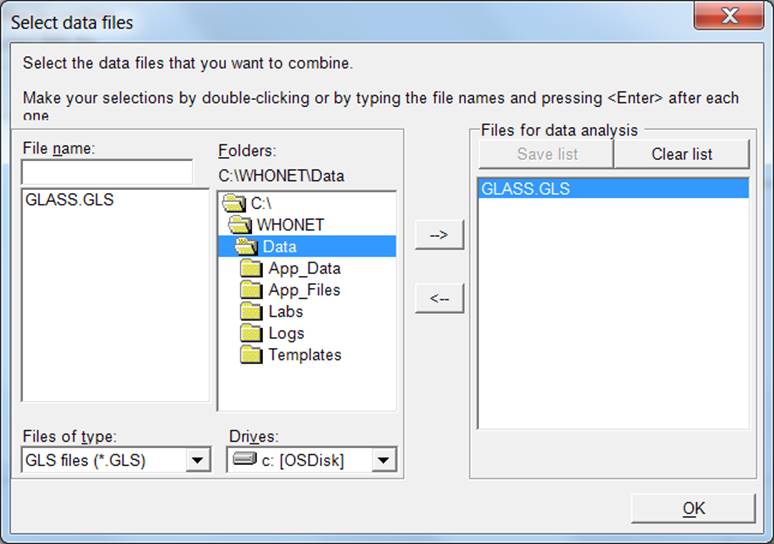
Ø Choose “All files (*.*)” to get a list of all files.
Ø Using the “-->” button, select the data files to be exported to the GLASS data exchange format.
Ø Click “OK”. You will return to the previous screen.
Ø Indicate a name for the new combined, export file.
Ø Under “Save as type”, change the export format from “WHONET” to “GLASS”.
WHONET will generate 2 files:
§ RIS file; and
§ Sample file.
Both files should be uploaded to the GLASS IT platform.[4]
The RIS file includes aggregated data submitted from all participating national surveillance sites on the number of resistant, intermediate and susceptible isolates detected in GLASS priority specimens and pathogens, stratified by gender, origin and age.
The sample file with “Sample statistics” includes the numbers of patients with specimens taken, stratified by the same variables as in the RIS file. That means the sample file will also include, for example, data on “negative” results provided these data were entered and are available in the source database (see also section 4.2, page 13, Pathogen).
By default, WHONET will save the new files in the C:\WHONET\Output folder named as following: GLASS-3 letter country code-data year-data set-file (RIS or Sample), though the user can change this if desired. For example:
1) RIS statistics: C:\WHONET\Output\GLASS-WHO-2015-DS1-RIS.txt
2) Sample statistics: C:\WHONET\Output\GLASS-WHO-2015-DS1-Sample.txt
Ø Ensure that the proposed WHONET “Data year” is the reporting year, otherwise change accordingly.
Note: Only data with the corresponding specimen date (year) and “data year” is included in the aggregated files. That means a specimen is included in the aggregated files, if specimen date is “01-01-2015” and if “data year” is “2015”; the specimen will be excluded in the aggregated files, if specimen date is “01-01-2015” and if “data year” is “2016”.
Ø “Data set” allows one to distinguish subsets of national aggregated data provided by a country where, for some reason, it is not possible to aggregate national data in the same way or when dividing the national data set has an important added value. This may be needed, for example, if the country has several different surveillance systems or there is a need to report data from different parts of the country separately. This may also be needed if, for example, the sample statistics (needed for generating a sample file) are missing in a large part of the country (see also the GLASS document “ A guide to preparing aggregated AMR data files” for more details). From the drop-down option, select the “Data set” code. Make sure it is the corresponding code for both RIS and sample file. The default option is “Data set 1” (code “DS1”).
The user can select the specimen types to include in the GLASS aggregated data file (per default all 4 GLASS specimens are included). If the user wishes to include only, for example, blood, make sure to choose blood for the RIS and the sample file.
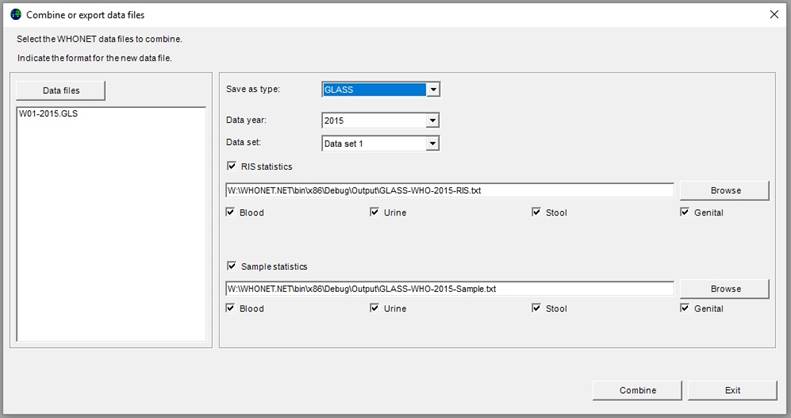
Ø Click on “Combine” to begin the export.
WHONET will read the selected data files and export all indicated specimen types and GLASS-requested pathogens and antimicrobial susceptibility interpretation results.
Ø After completion of the data export, WHONET will indicate the number of isolates included in the export file, followed by the Data Check Feedback report, as described in section 7.
6.3 WHONET rule for excluding “repeat isolates” in the GLASS exchange format
According to GLASS protocol, “repeat isolates” (“duplicates”) from a patient should be excluded. The Clinical and Laboratory Standards Institute (CLSI) document “M39 – Analysis and Presentation of Cumulative Antimicrobial Susceptibility Test Statistics” recommends that laboratories use the first isolate per species for the analysed time period and data subset when calculating susceptibility and resistance proportions for purposes of developing guidelines for empiric therapy. When generating the GLASS export file, WHONET automatically removes repeats following this recommendation – “first isolate per patient” regardless of susceptibility results. Consistent with the CLSI recommendation for data subsets, the WHONET export to GLASS will select the first isolate from each specimen type and “Infection origin”, e.g. E. coli first isolates from urine and from blood from the same patient will both be included in the GLASS export files.
7. GLASS Data Check and Feedback Report
Ø After completion of the data files export, the following screen will appear:
Ø By clicking on “Yes”, the following GLASS validation screen with selection options will open:
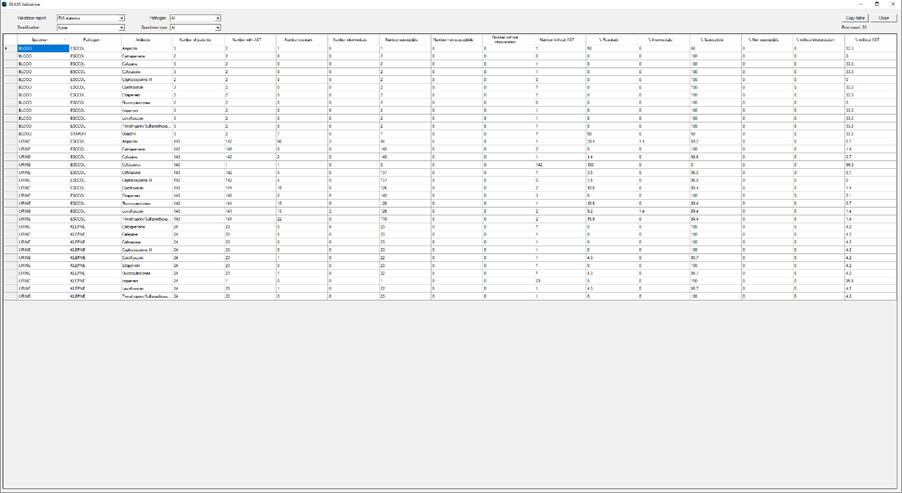
Ø The user can select between validating: 1) the “Sample file” (sample totals); and 2) the “RIS file” (RIS statistic). By default, the user is presented with the “Overall” statistics (all genders, age groups and origins aggregated together) or alternatively on the basis of each of the three stratification variables (“Gender”, “Age” (as age group) and “Origin”). The pathogen filter is not applicable to “Samples” validation. The user may also use the “Copy table” feature at any time to copy the visible table data. This is useful if one wishes to work with these data in a spreadsheet programme for example.
Validation of “Samples statistics”:
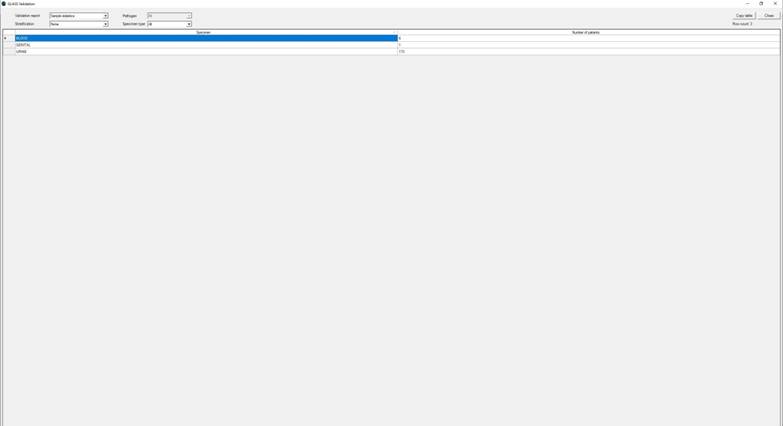
Ø The sample statistics display the total number of patients sampled for each specimen and included in the source data file according to the GLASS definitions.
The below screen grab displays the “Sample statistics” stratified by age group.
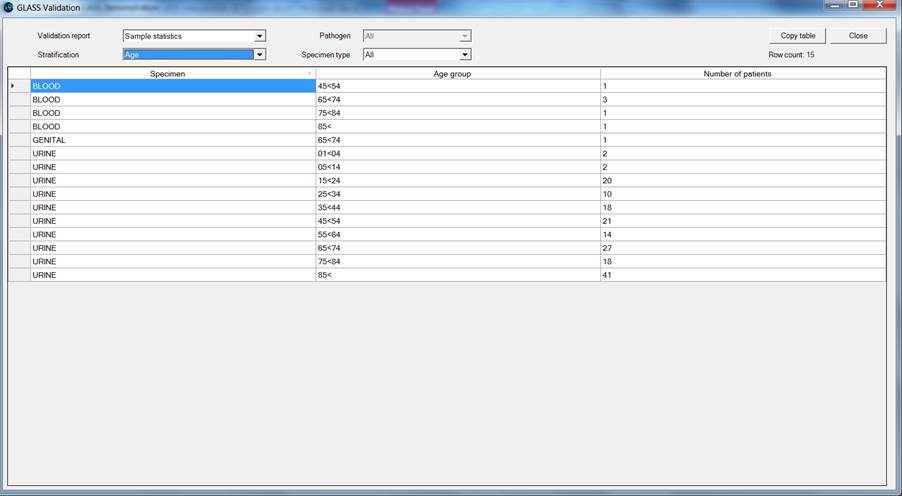
Validation of “RIS statistics” stratified by age group:
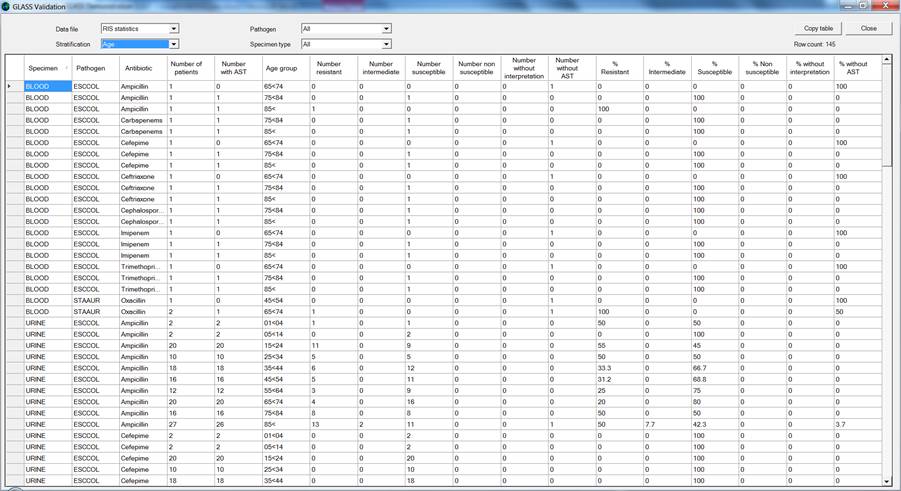
If any errors or unexpected findings are noted during the validation process, the user should review the original data files, edit if warranted, and then re-export the data.
8. Upload the GLASS-formatted data file to the GLASS IT platform
When data is exported using the steps described above, WHONET will create a GLASS “RIS file” (for example GLASS-WHO-2015-DS1-RIS.txt) and a GLASS “Sample file” (for example GLASS-WHO-2015-DS1-Sample.txt). By default, these files will be stored in the C:\WHONET\Output folder. GLASS instructions for uploading these two files to the GLASS IT platform are available.[5]
Annex 1 – Modify laboratory configuration
This annex provides further explanations on the WHONET module for modifying the laboratory configuration for antibiotics, locations and data fields. For more detailed information, please consult the relevant WHONET tutorials, available from http://whonet.org/documentation.html.
1. General laboratory information
Step 1: Collect information on antibiotics, test methods and breakpoints specific to the participating laboratory.
Step 2: Start WHONET by double-clicking on the WHONET icon. When the user enters “New Laboratory”, the “Laboratory configuration” screen appears.
This screen will also appear after the user has created a new GLASS laboratory following the steps in section 3.1. At the end of this process the user will be asked whether any additional modifications are required: select “Yes”.
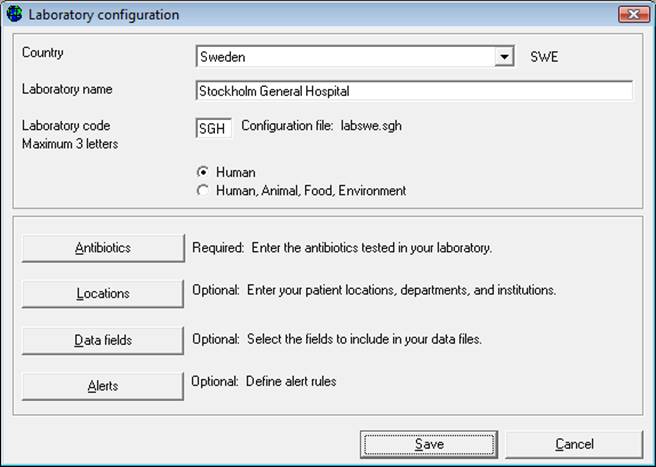
Step 3: Describe Laboratory (Required): Choose the country and enter the name of the laboratory. Enter a three-letter laboratory code.
Note: After entering this information WHONET will automatically create a corresponding laboratory configuration file name with the name labccc.xxx, where ccc = 3-letter ISO country code and xxx = 3 characters of laboratory code.
Step 4: Antibiotic Configuration (Required): Click on “Antibiotics” to configure the antibiotics. Enter the list of antibiotics that are being used by the laboratory. Click “OK” when finished to return to the main “Laboratory configuration” screen. For details see section 2 below: “Antibiotics”.
2. Antibiotics
To indicate which antibiotics, methods and breakpoints are being used in the participating laboratory the user must enter “Antibiotic configuration”.
Note: During data entry the software performs automatic interpretation based on the breakpoints entered in this configuration.
Instructions
Step 1: From the main Laboratory Configuration screen, click on “Antibiotics”: the screen below appears. The “WHONET antibiotic list” appears on the left, while the antibiotics that the user may want to select will appear on the right under “Local antibiotic list”. The list on the right will already contain GLASS antibiotics if the user has chosen the GLASS laboratory configuration.
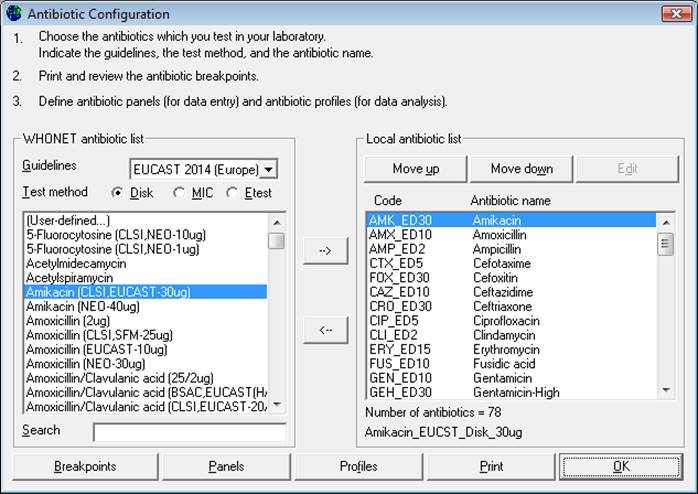
Step 2: For each combination of antibiotic, method and testing guidelines:
§ Select the correct testing guidelines (CLSI, EUCAST, etc.); and
§ click on the correct test method (disk diffusion, MIC, ETestÒ); and
§ select the correct antibiotic (and disk potency for disk diffusion testing).
An antibiotic can be selected by double-clicking on it or, alternatively, by clicking on it once and then clicking the right arrow button “-->”.
Once all the antibiotics have been entered, review the list and make any needed corrections. To remove an antibiotic from the local antibiotic list, double-click on the antibiotic or single-click on the antibiotic and hit the left arrow button “<--“. The sequence of the antibiotics may be changed using the “Move up” and “Move down” buttons.
Note: Each antibiotic test is given a code (up to nine letters) consisting of: the three-letter antibiotic code, a one-letter code indicating the guideline reference (e.g. N=CLSI, formerly NCCLS, E=EUCAST), a one-letter code indicating the test method (D=disk diffusion, M=MIC, E=ETestÒ), and a disk potency for drugs tested by disk diffusion. For MIC or ETestÒ it is only necessary to select the correct antibiotic and correct guideline – the disk potency indicated is not relevant. For example, the code GEN_ND10 represents: gentamicin, CLSI (NCCLS), disk diffusion, 10 µg, while GEN_EM is: gentamicin, EUCAST, MIC.
Step 3: When satisfied with the list, the user can choose from the following options:
§ “OK” to return to main “Laboratory configuration” screen. Note the information has to be saved by clicking on “Save” in the main “Laboratory configuration” screen.
§ “Panels” and “Profiles” are optional: to facilitate data entry, the user may wish to use “Panels” to indicate which antibiotics are tested for each kind of organism. For details refer to the complete WHONET manual.
§ “Print”: obtain a printout of the antibiotics and their breakpoints. It is recommended that breakpoints are printed for review and reference before data entry commences.
§ “Breakpoints”: to review, modify and update the antibiotic breakpoints. See next section 3 “Antibiotic breakpoints”.
3. Antibiotic breakpoints
WHONET will automatically load the most recent breakpoints for the antibiotics that have been indicated. Although not recommended for most users, it may happen that a laboratory uses other breakpoints then standard reference breakpoints, and for these situations it is possible to edit the WHONET-provided standard breakpoints.
Instructions
Step 1: From the “Antibiotic configuration” screen, click on “Breakpoints”. The following screen will appear.
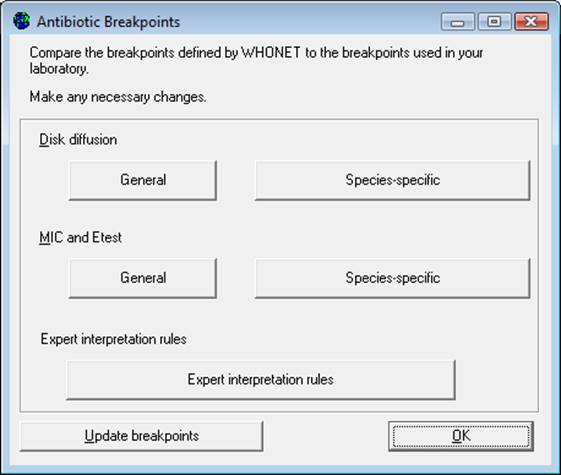
Step 2: The following options are available:
§ “General”: review and edit the list of general disk diffusion or MIC breakpoints. For details, see Step 3a.
§ “Species-specific”: review and edit the list of species-specific disk diffusion or MIC breakpoints. For details, see Step 3b.
“Expert interpretation rules”: These are not required for GLASS data management. Refer to the complete WHONET manual for further details.
“Update breakpoints”: The WHONET antibiotic definition files are updated annually as new recommendations from the reference authorities become available. If the user downloads WHONET on an annual basis, the download will include the new breakpoints. WHONET will not automatically use these new breakpoints for an existing antibiotic list until the user clicks on “Update breakpoints” to replace the breakpoints that have been set for the laboratory with the antibiotic breakpoints to be found in the most recent antibiotic definition files.
“OK”: When finished reviewing or modifying the antibiotic breakpoints, selecting “OK” will return the user to the “Antibiotic configuration” screen.
This example illustrates the procedure:
Step 3a
Editing General breakpoints: Click on Disk diffusion “General” breakpoints: the screen below appears. (If the option for MIC and ETestÒ breakpoints is selected, the same type of screen will appear.)
Note: Since 2013, WHONET no longer defines “General” breakpoints for EUCAST. Instead all breakpoints are associated with specific species and can be found under “Species-specific breakpoints”.
Click on any breakpoint values that are to be changed. The relevant values may be edited under “R”, “I”, or “S”.
Note: If the user changes a value e.g. for “I”, the other values change accordingly.
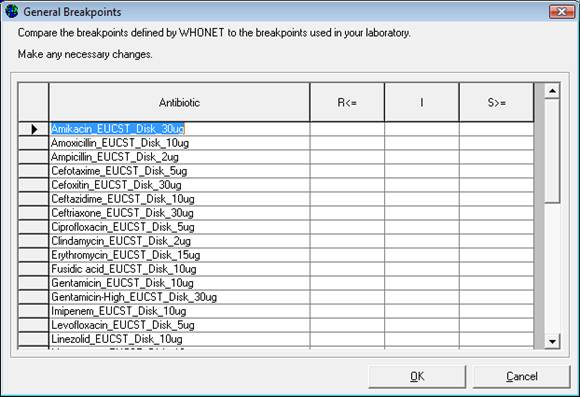
Once there are no more changes to be made, click “OK” to return to the previous screen.
Note: When entering the MIC breakpoints for antibiotic combinations, such as trimethoprim/sulfamethoxazole, enter the concentration of the first constituent. These dilutions usually follow the standard 1, 2, 4 µg/ml, etc. series.
Step 3b
Editing Species-Specific Antibiotic Breakpoints: A screen similar to the one below will appear. In most countries, changes to these WHONET-provided reference breakpoints are not required. However, if the user wishes to make modifications to the breakpoints, changes to the table breakpoints below may be made using the same procedure as described in Step 3a.
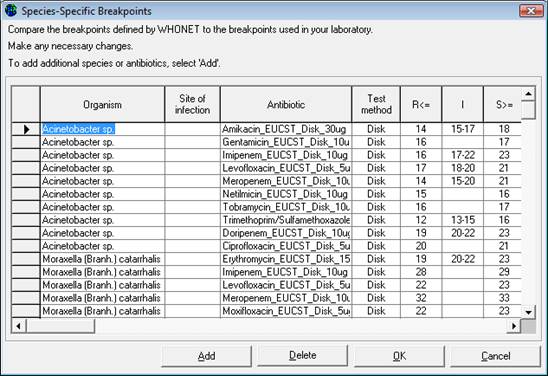
Adding Species-Specific Antibiotic Breakpoints: If it is necessary to enter additional species-specific breakpoints, select “Add”, and the screen below will appear. Indicate the organism-antibiotic combination for which one wants to define breakpoints by clicking on the desired organism and the desired antibiotic. When finished, select “OK” to return to the previous screen where the breakpoints may be entered.
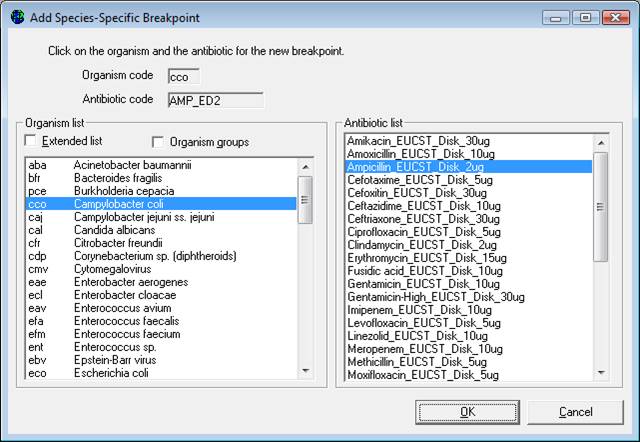
Deleting Species-Specific Antibiotic Breakpoints: To delete a species-specific breakpoint, click on the relevant row of the table, and click ‘Delete’.
After completing changes to the breakpoints, select “OK” to return to the “Antibiotic Breakpoint” screen. Select “OK” again to return to the “Antibiotic Configuration” screen.
4. Locations
This option from the main screen “Laboratory configuration” is optional, but is of value if the user would like to keep track of patient locations and medical services from which samples are taken. A detailed description is found in the main WHONET manual.
5. Modifying data field configuration
A set of “standard” data fields is defined automatically by WHONET. “Standard” fields include routine information such as: unique patient ID, location type, specimen date, specimen type, organism/pathogen, antibiotic panel, etc. To combine additional surveillance activities with the GLASS data collection, the user may want to add some “additional” data fields: see instructions below.
Instructions
Step 1: From the main “Laboratory configuration” screen, select “Data fields” and click on “Modify list”. A screen similar to the one below should appear. The lists of WHONET data categories (clinical, infection control, etc.) and data fields (diagnosis, admission date, etc.) from which the user may select appear on the left. If the user selects “GLASS”, this will show the GLASS data fields in the box “Data fields”. Select all and click the right arrow. They will be added to the list of data fields that appear to the right (also containing the standard data fields).
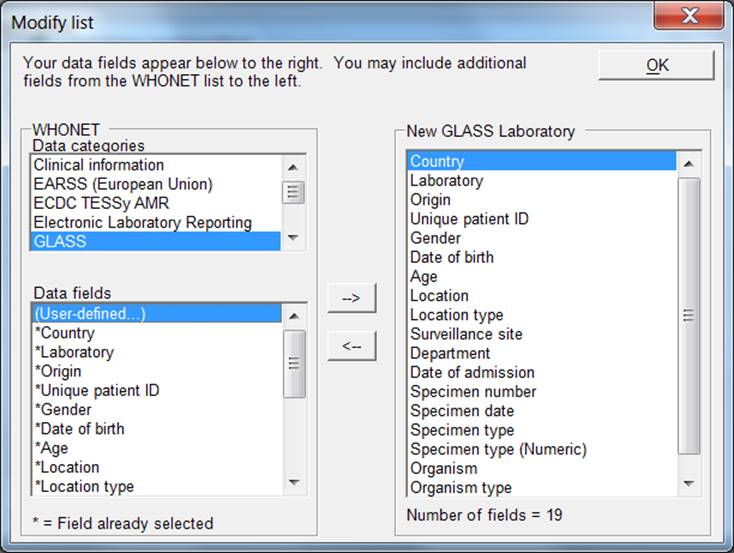
Step 2: If the user would like to include more additional fields in their data files, then they may be selected here also.
Note: By default, WHONET will check the validity of GLASS codes entered during data entry.
Step 3: Leave this screen by clicking on “OK” twice and click on “Save” to save this laboratory configuration.
Annex 2 – BacLink introduction
Many laboratories around the world already have well-established computer databases that meet the day-to-day needs of clinical reporting, specimen processing and long-term data storage. Unfortunately, most of these systems have limited capacity for sophisticated data analysis. It is in these areas that WHONET is a valuable supplement to existing systems.
The purpose of the BacLink software is the conversion and standardization of microbiological data from existing systems into WHONET. Data can be converted on a weekly, monthly, or ad hoc basis: In a number of institutions, it has been possible to automate and schedule the entire process.
In using BacLink to convert data to the WHONET data format, laboratories have the benefits of:
§ flexible data analysis capabilities; and
§ the ability to share standardized data with other laboratories, for example in a national surveillance network.
BacLink can import data with a variety of data structures including:
§ Simple data structures: delimited text files (for example created from Microsoft Excel), or Microsoft Access, EpiInfo, or dBASE files;
§ Commercial organism and antimicrobial susceptibility test instruments, such as Vitek, Microscan, or Phoenix; and
§ Laboratory information systems (LIS): These may be commercial systems, such as Cerner or Meditech, or developed in-house by the facility IT staff. Most such systems have the ability to create simple delimited text files that can be imported by BacLink.
Further details on the steps for configuring BacLink and data capture are provided in a series of BacLink tutorials beginning with BacLink 1 – Getting started that can be downloaded from the WHONET Home Page, http://www.whonet.org/documentation.html.