`
WHONET
Cluster Detection with SaTScan™
WHO Collaborating Centre for
Surveillance of Antimicrobial Resistance
Boston, Massachusetts
June 2006
WHONET Tutorial – Cluster Detection with SaTScan™
For many years, WHONET has had the ability to present temporal trends in organism frequencies and resistance proportions in the form of descriptive statistics and graphs. The user could then examine these trends and graphs individually in efforts to detect and characterize possible community or hospital outbreaks of microorganisms. WHONET was not able to highlight potential clusters in an automatic fashion in order to focus the attention of the software user on possible outbreaks or to provide statistical guidance as to whether observed trends were statistically significant.
To facilitate the early and broad detection of possible outbreaks, we have integrated a powerful freeware tool developed for purposes of cluster detection in public health data.
SaTScan™, a trademark of Martin Kulldorf, was developed under the joint auspices of Martin Kulldorf, of the National Cancer Institute and of Farzad Mostarashi at the New York City Department of Health and Mental Hygiene. Dr. Kulldorf is an Associate Professor and Biostatistician at Harvard Medical School and Harvard Pilgrim Health Care, Department of Ambulatory Care and Prevention, Boston, USA.
Kulldorf M. and Information Management Services, Inc. SaTScan™ v.7.0: Software for the spatial and time-scan statistics. http://www.satscan.org, 2006.
The software permits a number of algorithms for the detection of event clusters. Options include retrospective or prospective cluster detection; purely temporal, pure spatial, or space-time clusters; and flexible parameter selection for space and time variables. In this first version of an integrated WHONET-SaTScan package, WHONET is using a space-time permutation probability model, using Monte Carlo simulations. In collaboration with Dr. Kulldorf in a five-year NIH project entitled “Modeling Infectious Disease Agent Study” (MIDAS), we will test out a number of additional algorithms, models, and parameters, and these optimized routines will eventually be offered through the WHONET user interface.
In WHONET 5.4, SaTScan statistics have been integrated into two standard analysis features: 1. Isolate listing summaries and 2. Resistance profile summaries. Two new analysis options specific to SaTScan have also been added: 1. SaTScan – Cluster detection in Data Analysis and 2. SaTScan – Space-and-Time cluster detection in Quick Analysis.
Part 1. SaTScan and Isolate listing summaries
Part 2. SaTScan and Resistance profile summaries
Part 3. SaTScan – Cluster detection (Data Analysis)
Part 4. SaTScan – Space-and-Time cluster detection (Quick Analysis)
Part 1. SaTScan and Isolate listing summaries
Open the WHO Test Hospital, and go to Data Analysis. Go to Analysis Type, and select Isolate listing and summary. At the bottom of the screen, click on the option “Include SaTScan alerts”. By doing this, the SaTScan statistics will be integrated directly into the standard Isolate Listing Summary output.
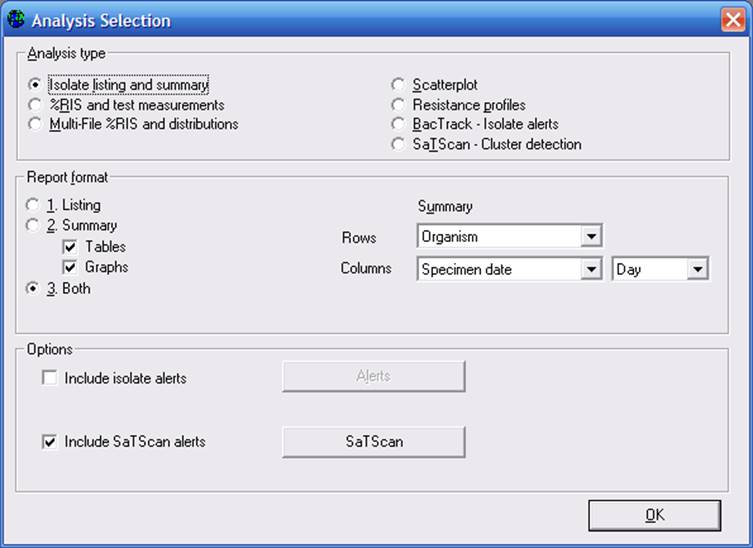
Note: In the above example, the column variable has been set to “Specimen date” by “Day”. In most cases, you should probably leave this option set to “Month” or “Week”. But for purposes of this tutorial, we have set the option to “Day” because the sample dataset only includes one month of data.
To see the details of the SaTScan analysis, click on “SaTScan” to see the below screen.
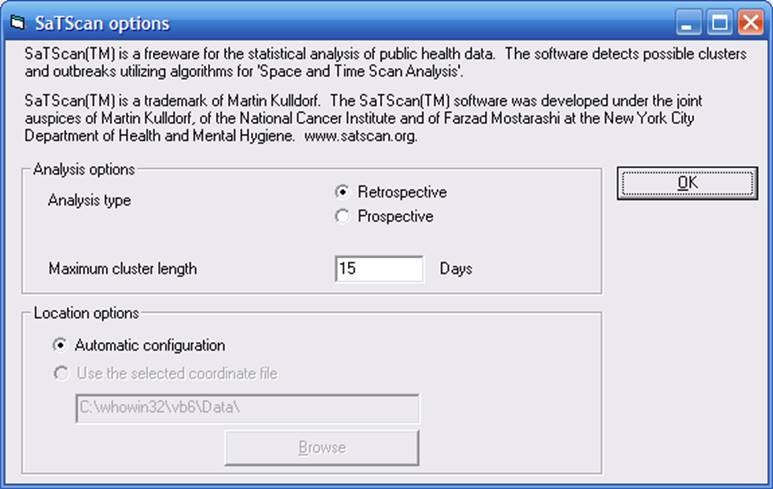
On this screen, you will see a small minority of all of the parameter options available in SaTScan. The use of SaTScan to study microbiological laboratory data and hospital infections data is very new, and will be a major focus of the 5-year NIH grant. As we gain more experience with the use of SaTScan for microbiology data, the number of options included in WHONET will increased and the suggested default parameter values will change to reflect optimal use of the program by microbiologists, infection control, and clinical staff.
In the above example, the user has access to two SaTScan variables.
Analysis type: Retrospective vs. Prospective. Retrospective analysis looks for infectious disease clusters at any time during the data analysis time period – both old clusters and new clusters. This option would be very useful in a retrospective look at available data for any possible outbreak. Prospective analysis only looks for clusters that potentially are still ongoing. This option is appropriate for the prospective detection of outbreaks in the new data.
Maximum cluster length: Indicate the number of days for the greatest length of a possible outbreak. At present, the default is set to 15 days. In other words, SaTScan will look for clusters that are anywhere from 1 day to 15 days in length. If an outbreak is longer than this period, SaTScan possibly may not detect it or possibly may detect only part of it. To find the greatest possible number of potential clusters, you can put a large value for the Maximum cluster length. A long period can slow SaTScan down considerably, so you can experiment to see what impact this value has on 1) the number of detected clusters and 2) the time period needed for the analysis.
Note: In SaTScan, the largest possible value of the Maximum cluster length is half the analysis time period. For example, if you have one year (365 days) of data to scan, the largest value that SaTScan will accept is 182 days. It is not a problem if you enter a higher value into WHONET. WHONET will dynamically adjust the value before it sends the analysis parameters to SaTScan.
Location options: In WHONET so far, we are not yet taking advantage of SaTScan’s ability to handle geographic data. For outpatients, such variables could include zip code or street address GPS latitude and longitude. For inpatients, it is possible describe to SaTScan the geographic (hospital physical location) or functional (clinical care service) relationship between patient care units. So at the present time, WHONET does an automatic configuration of patient locations, treating each ward as a separate “island” of care, unconnected to any other wards.
In the future, it will be possible for WHONET users to describe the ward structure of hospital units in a SaTScan “coordinate file” in a simple user interface. If a user does take the time to describe the geography of their hospital to WHONET, this option will permit the user to select the coordinated file created.
After you select the desired features, click “OK” to return to the main analysis screen.
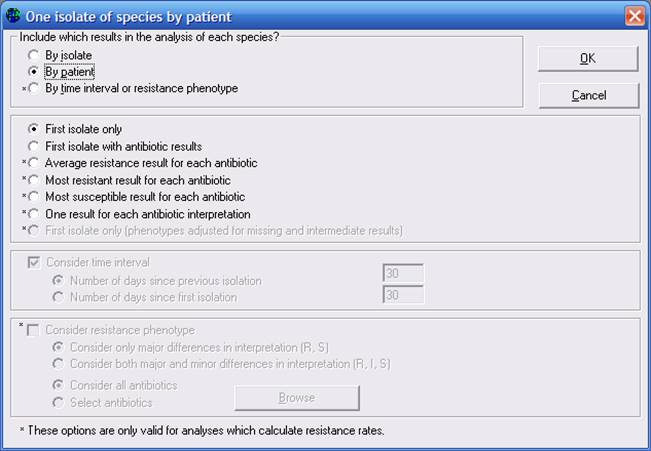
Then click on “One per patient”. Select the options “By patient”, “First isolate only”.
Note: It would be probably more useful for cluster detection to put the option “By time interval or resistance phenotype”, for example with a 30 day window between isolates. This option does exist for %RIS calculation, but unfortunately, this option is not yet ready for other analyses, such as isolate listings and resistance profiles. This will be addressed in the near future.
Note: We do not yet know what criteria would be most effective in detecting real infectious disease outbreaks, and whether the criteria should differ for different organisms. SaTScan does an excellent job of finding statistically defined “clusters” of events, but the clinical interpretation of these clusters will depend on the experience and insight of infection control and clinical staff. With more testing on real datasets, we will aim to provide tested and validated recommendations for setting the cluster detection parameters.
Finally, click “OK” and Begin Analysis. First you will see the standard isolate listing. (SaTScan makes no changes to the isolate listing. If you do not wish to see the listing, you should choose “Summary” as the Report format rather than “Both”.)
When you click Begin Analysis, WHONET will read the data files, and prepare the files needed by SaTScan to operate. WHONET will then send this information to SaTScan. A small icon will appear at the bottom of the screen to tell you that SaTScan is running in the background. When SaTScan is finished, this icon will disappear automatically.
After SaTScan is finished analyzing the data, WHONET will read the SaTScan results, and integrate the statistics into the normal WHONET output, as in the below screen.
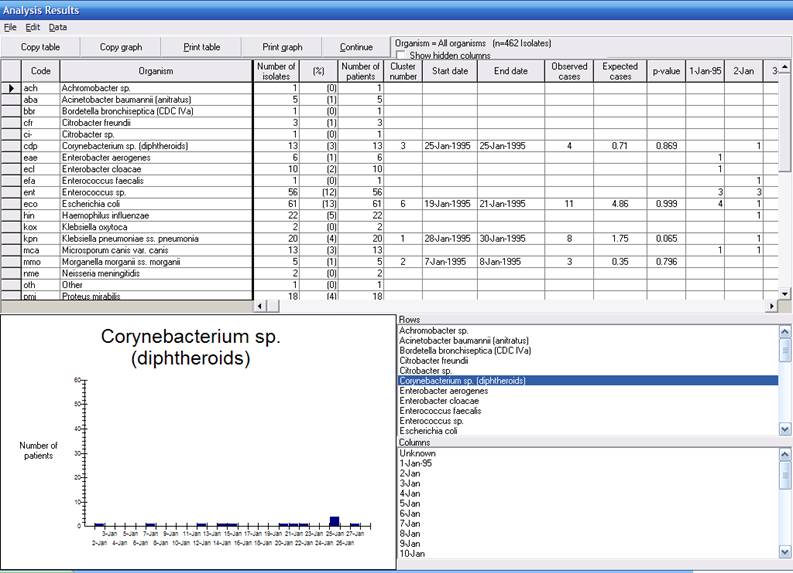
In addition to the typical columns for Isolate listing summaries, you see several additional columns in this output related to the clusters detected by SaTScan.
In the sample dataset with one month of data, WHONET found 39 bacterial species. SaTScan identified possible clusters in 7 of these species. Cluster number 1 is the cluster of greatest statistical significance (smallest p-value), etc. Only one of the identified clusters has a small p-value:
Organism Klebsiella pneumoniae
Cluster dates January 28-30
Observed 8 patients
Expected 1.75 patients
p-value 0.065
This particular
organism is depicted in the below graph, confirming that there does indeed seem
to be a rise in the number of patients with K. pneumoniae at the end of
the month.
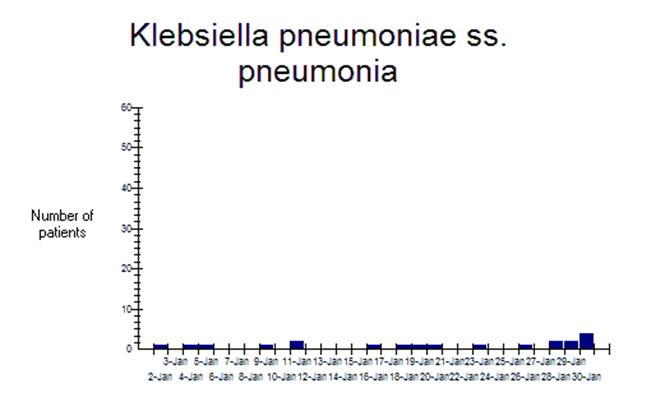
Part 2. SaTScan and Resistance profile summaries
In Part 1, we saw how SaTScan statistics can be applied to organism frequency data. In this section, we will see how the same statistics can be applied to resistance profile data. For the output columns, put “Specimen date” by “Day”. (As above, we are selecting “Day” in this tutorial because there is only one month of data.)
Go to “Analysis Type”, and select “Resistance profiles”. Check on the box for “Include SaTScan alerts”.
For “One per patient”, put “By patient”, “First isolate only”. For “Organisms”, select S. aureus. Then begin the analysis. WHONET will first show you the Resistance profile listing. This is not affected by the SaTScan analysis. Hit Continue to proceed to the Resistance profile summary.
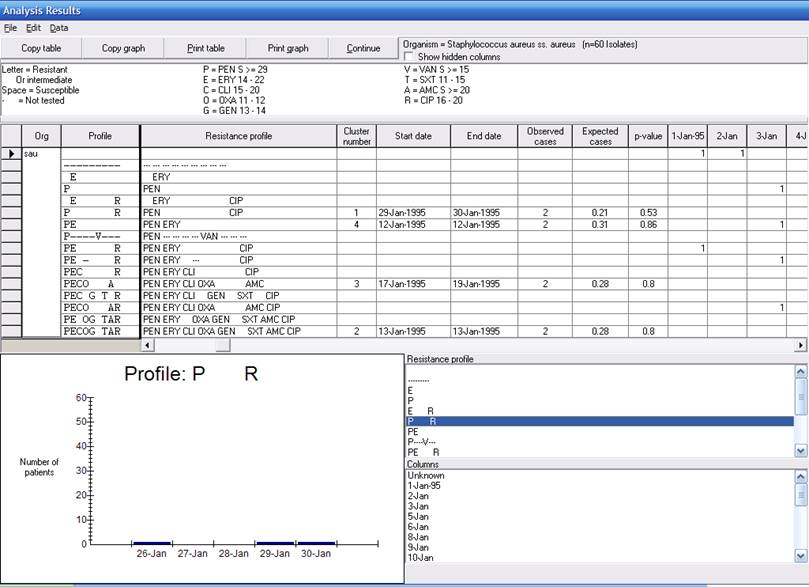
WHONET found 60 isolates of S. aureus, which cluster into 16 different resistance phenotypes. (The results “-” and “---” indicated that the indicated antimicrobials were not tested). From the 16 resistance phenotypes, SaTScan identified 4 possible clusters. Cluster number 1 has the following characteristics.
Resistance profile PEN CIP (non-susceptible only to PEN and CIP)
Cluster dates January 29-30
Observed 2 patients
Expected 0.21 patients
p-value 0.53
The graph for this resistance profile is given above. Even though the number of isolates is very small, and the p-value is not, this cluster may still be of interest. In the entire month of January, this resistance profile was only seen three times, all at the end of month – once on January 26 (not detected by SaTScan), January 29, and January 30. In a further investigation, it would be useful to look at the patient locations. If you refer back to the Resistance profile listing, you will notice that the three isolates came respectively from location “xx = Unknown”, “er = Emergency Room”, and “op = Outpatient”. So there is no epidemiologic link between these three patients with the available data, but one possibly would be revealed with further investigation of the patient medical records.
Part 3. SaTScan – Cluster detection (Data Analysis)
In parts 1 and 2, we saw how SaTScan results can be integrated with standard WHONET analyses. In the next two parts, we will focus only on the information generated by SaTScan.
In “Analysis type”, select “SaTScan – Cluster detection”. At the bottom of the screen, click on “SaTScan” to see additional options. In all above analyses, we used a Retrospective analysis. For this example, choose Prospective analysis.
For organisms, put “All organisms”. For One per patient, put “By patient”, “First isolate only”. Begin the analysis.
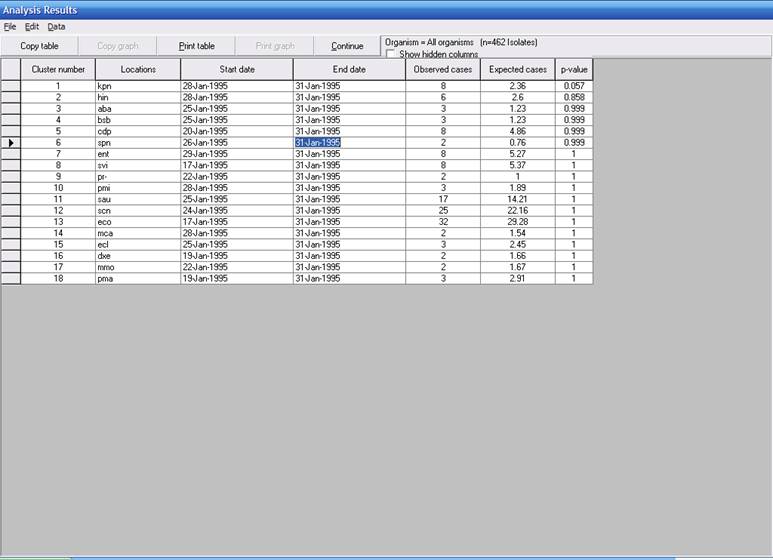
In Part 1, we saw that January had 39 bacterial species. Of the 39 species, SaTScan has identified 18 possible “ongoing” clusters, in other words clusters that potentially could include January 31, the final date of the analysis period. Of the 18 clusters, only one has a small p-value
Organism Klebsiella pneumoniae
Cluster dates January 28-31
Observed 8 patients
Expected 2.36 patients
p-value 0.057
This is the same cluster detected in Part 1 of this tutorial.
Part 4. SaTScan – Space-and-Time cluster detection (Quick Analysis)
Now exit from the usual Data analysis program, and select “Quick analysis”.
Choose the option “SaTScan – Space-and-Time cluster detection”. If you click on “Edit”, you will see the same SaTScan options menu which we saw in Data analysis.
For Data files, choose the w0195who.tst data file. Click on Begin analysis.
WHONET will display for you a two-part output. Part A – SaTScan report is the actual report prepared by the SaTScan software summarizing the details of the analysis performed and all statistical findings. This output begins with a brief summary of the analysis performed and the data summarized.
It then continues with a list of all of the identified clusters in order of decreasing statistical significance. Finally at the bottom of the output, you will see the detailed parameter settings used for the analysis.
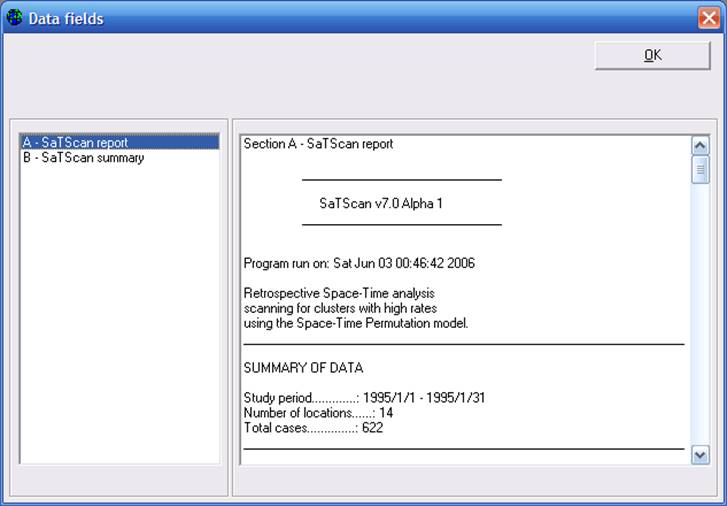
If you continue with Part B – SaTScan summary, you will see that WHONET has extracted the most important pieces of information for display to the user in a concise format. (Note: in this first version of WHONET 5.4, the formatting of the Quick Analysis results has not been optimized. This will be improved in the near future.)