WHONET
Data analysis 2
WHO Collaborating Centre for
Surveillance of Antimicrobial Resistance
Boston, Massachusetts
June 2006
WHONET Tutorial – Data Analysis 2
This tutorial covers the following topics.
Part 1. Getting started and review
Part 2. One-per-patient options
Part 3. Isolate selection criteria
Part 4. Other options
Part 5. Scatterplots
Part 6. Antimicrobial resistance profiles
Part 7. What next?
The tutorial Data Analysis 1 introduced the WHONET data analysis program and illustrated some of the most common analyses performed with WHONET: 1. %RIS and test measurements; and 2. isolate listings and summaries. In this tutorial, you will see how to select patients or isolates that meet certain criteria, options for handling multiple organism isolates per patient, and two additional analysis features.
For information on the following analysis options, refer to the appropriate tutorials:
Expert system
Macros and Excel reports.doc
Cluster detection with SaTScan
Part 1. Getting started and review
If WHONET is not currently running, double-click on the WHONET icon on your desktop to begin the software. From the list of available laboratories, select the laboratory called “WHO Test Hospital”, and click on “Open laboratory”. From the main WHONET screen, click on “Data analysis” and “Data analysis” again. You will now see the main WHONET analysis screen.
For Analysis type, select “%RIS and test measurement” and for Report Format, also choose “%RIS and test measurement.
For Organism, select S. aureus = sau.
For Isolates, there should be no selection criteria defined.
For Data Files, select w0195who.tst.
If you have done these steps properly, you should see the following screen.
Figure 1. %RIS and test measurements with S. aureus.
Then click on “Begin Analysis” to get the results. On the output screen, notice that oxacillin was tested 85 times, and 10.6% of the isolates were resistant. After reviewing the rest of the results, click on “Continue” to return to the main analysis screen.
Part 2. One-per-patient options
In the preceding analysis, WHONET included results from all S. aureus isolates in the analysis (except for laboratory and quality control isolates). For some analyses, such as isolate listings, including results from all isolates is appropriate.
However, for calculations of %Resistant or %Susceptible, it is usually recommended to make an adjustment to account for the fact that some patients have multiple culture results for the analyzed species. Patients who remain a long time in the hospital or patients with complicated clinical courses may have multiple isolates of the species in question, but outpatients and patients with simple clinical histories may only have a single isolation. If no adjustment is made to the calculation, the statistics will be biased towards the results of the patients with multiple isolates, frequently a subset with higher rates of resistance than other patients.
The U.S. Clinical and Laboratory Standards Institute (CLSI) has published a document “M39 – Analysis and Presentation of Cumulative Antimicrobial Susceptibility Test Statistics”, and this document recommends that laboratories use the first isolate per species for the analyzed time period when calculating susceptibility and resistance proportions for purposes of developing guidelines for empiric therapy.
WHONET offers several strategies for handling multiple patient isolates, including the CLSI recommendation of “first isolate per patient”. To see these options, click on “One-per-patient”. At the top of the screen, choose “By patient”. Then choose “First isolate only”.
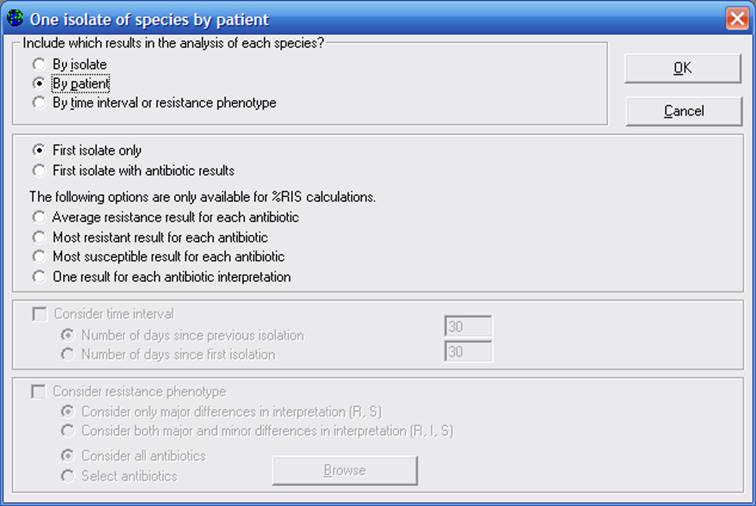
Figure 2. One-per-patient options. The CLSI recommendation is to use the first isolate per species in the analyzed time period.
Click “OK”, and then “Begin analysis” to get the following results. You will notice now that WHONET is displaying results of 57 patients with S. aureus tested for oxacillin, and 14.0% of the first isolates were resistant to oxacillin (In most analyses of hospital pathogens, the “by isolate” approach wield yield a higher estimate of %resistance than the “by patient” approaches, but at this example demonstrates, this is not always the case.)
When finished reviewing the other results, click “Continue” again to return to the main analysis screen.
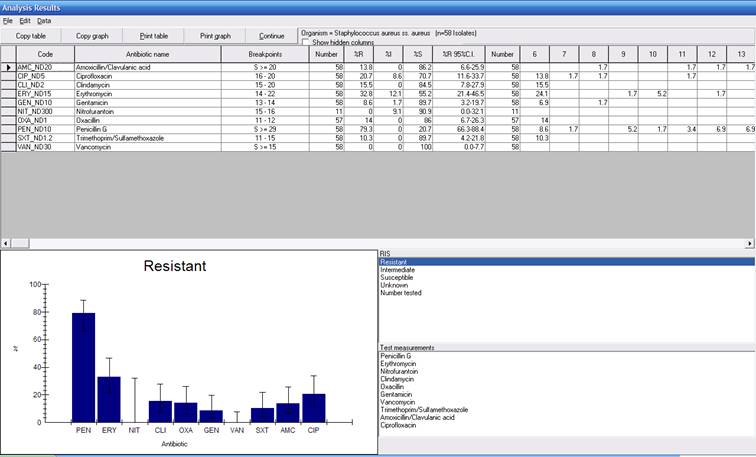
Figure 3. %RIS for S. aureus using the first isolate of S. aureus per patient.
After you return to the main screen, click “One-per-patient”, and set the option back to “By isolate”, and click “OK” to return to the main analysis screen.
Part 3. Isolate selection criteria
In the above analyses we analyzed all clinical isolates of S. aureus. (By default, WHONET automatically excludes quality control and other laboratory isolates). In many cases, users would like to select a subset of isolates or patients that meet certain criteria: for example blood isolates from the neonatal intensive care unit or imipenem-resistant Enterobacteriaceae.
To define selection criteria, click on “Isolates”. You may choose among any of the patient, location, sample, or microbiological/antibiotic fields available.
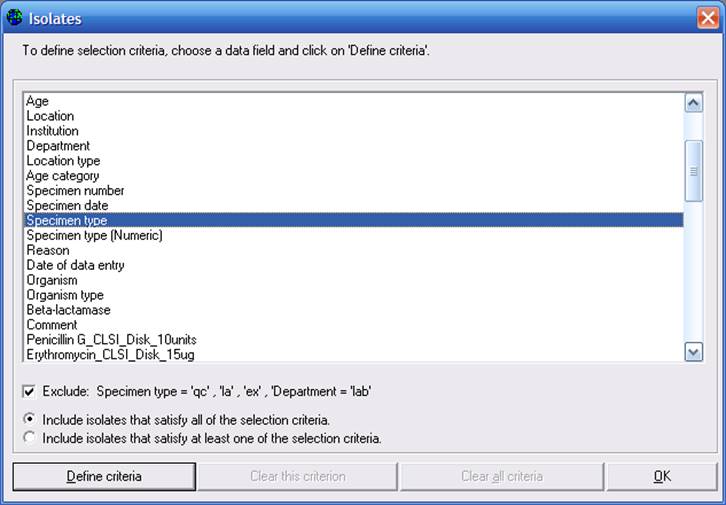
Figure 4. Isolate selection criteria. The “Specimen type” field is selected.
For example, double-click on “Specimen type” (or single-click on “Specimen type” and then click on “Define criteria”). Double-click on “Blood” to select this option, and click “OK”.
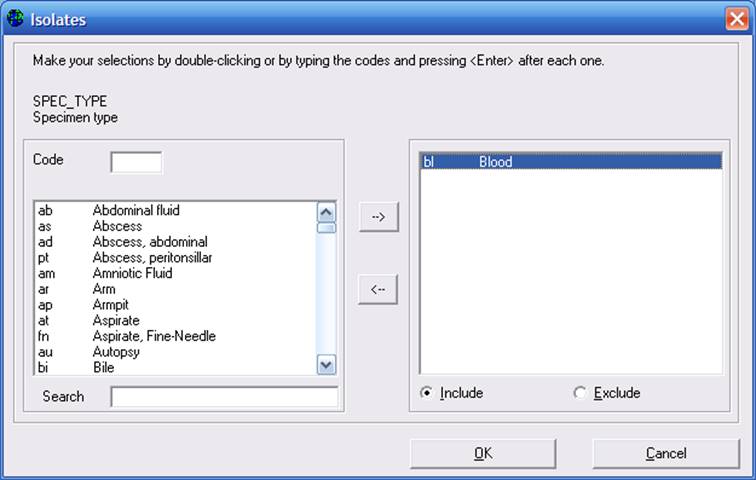
Figure 5. Defining selection criteria for “Specimen type”. You can “Include” or “Exclude” isolates with the criteria defined.
Now find “Oxacillin-Disk” on the list, and double-click. Choose “Resistant”, and “OK”.
There are a few other options on this screen that you should leave unchanged for this tutorial:
Exclude laboratory samples: To avoid mixing clinical from non-clinical isolates in your institution’s summary statistics, WHONET by default does not include the isolates in which the specimen type is equal to “qc” (quality control), “la” (laboratory sample), or “ex” (external quality control) or if the department is equal to “lab”. If you would like to include these results in your analyses, remove the check in this box.
Include isolates that satisfy all of the selection criteria / Include isolates that satisfy at least one of the selection criteria: If you choose the first of these options, WHONET will search for isolates that meet all of the selection criteria. In this example, WHONET will find results in which the isolate is from blood and is also resistant to oxacillin.
If you choose the second option, WHONET will search for isolates that meet at least one of the criteria. This would not be very useful in this example, but a more interesting example would be to ask for Enterobacteriaceae that are resistant to cefotaxime or to ceftazidime (or to both agents).
Delete this criteria / Delete all criteria. If you select these options, you can remove the selection criteria that you have selected.
After selecting the criteria “Specimen type = Blood” and “Erythromycin= Resistant”, click on “OK” to return to the main analysis screen. (Generally, it would be more interesting to study oxacillin-resistant strains than erythromycin-resistant ones, but in this small data set, there were no blood isolates of MRSA isolates.)
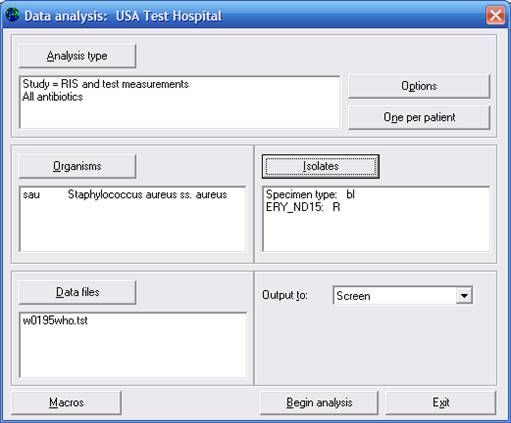
Figure 6. %RIS and test measurements for S. aureus with two isolate criteria defined: Specimen type = Blood and Erythromycin-Disk = Resistant.
Click on “Begin analysis” to repeat the %RIS analysis performed earlier, but using the selection criteria indicated here. The new output will show the %RIS statistics for blood isolates of erythromycin-resistant S. aureus (8 isolates in the small dataset). When you finish reviewing the results, click “Continue” to return to the main analysis screen.
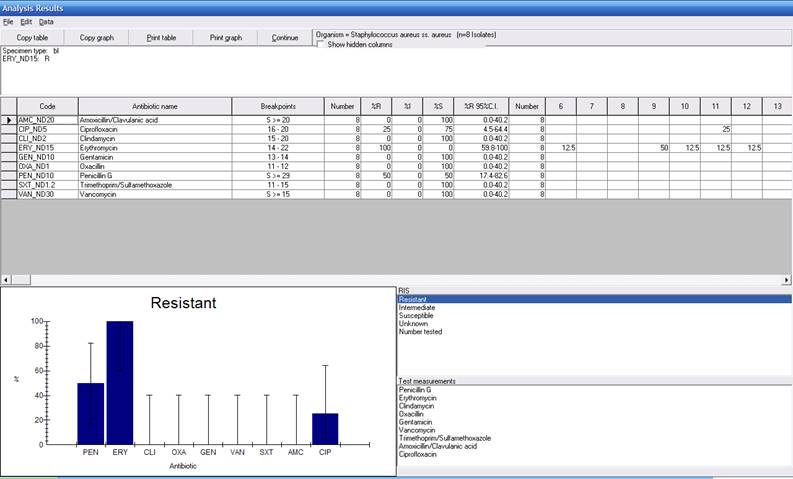
Figure 6. %RIS and test measurement results for S. aureus with two isolate criteria defined: Specimen type = Blood and Erythromycin-Disk = Resistant.
Before you leave this section, remove the selection criteria that you defined. To do this, click on “Isolates” and “Delete all criteria”. You will be asked to confirm your choice, so answer “Yes”.
Part 4. Other options
There are several additional options which permit small adjustments in the presentation of the results. To see these, click on “Options”, and review the options available to you. For this tutorial, make the following small change to the %RIS analysis.
For “%RIS and histograms”, change the option “Percent of isolates” to “Number of isolates”. Then click on “OK”, and “Begin Analysis”. In the output, you will see the number of isolates resistant, intermediate, and susceptible, rather than the percentages seen in the earlier analysis.
Click on “OK” to proceed to return to the main analysis screen.
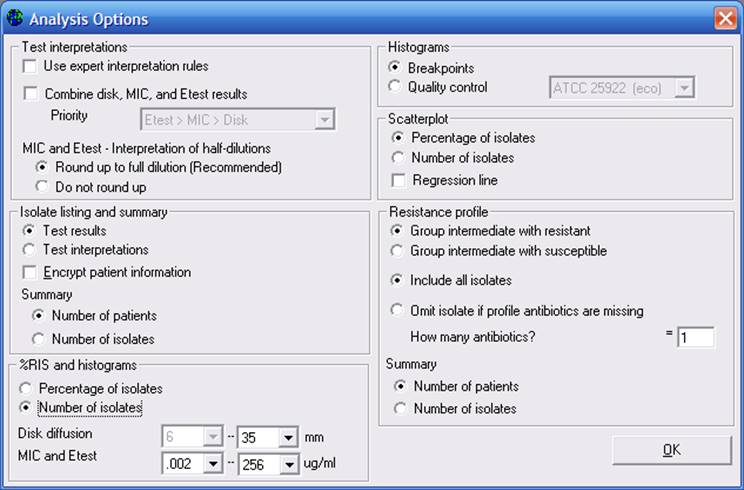
Figure 6. Additional analysis options.
Part 5. Scatterplots
In Data Analysis 1, you saw how to do isolate listings, summaries, and %RIS statistics. In this section, you will see how to compare the results of two antibiotic tests to each other.
Click on “Analysis type”, and select “Scatterplot”. In the lower part of the screen, you will select to antibiotic tests to compare. For the first example, put the Penicillin-Disk test on the X axis and the Oxacillin-Disk test on the Y axis. Then click “OK” and “Begin analysis” to get the following graph.
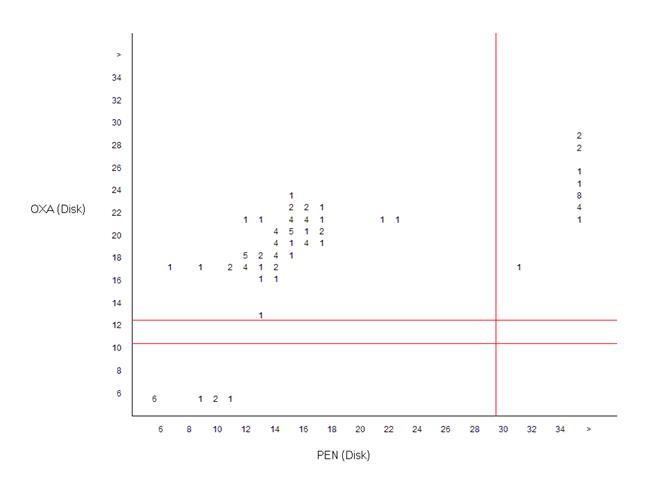
This graph shows the zone diameter distribution of isolates tested against both penicillin and oxacillin. Numbers in the figure represent percentage of isolates, and the red lines indicate the interpretative breakpoints.
For example, in the upper right-hand corner of the graph, there are a number of isolates with large zone diameters for penicillin (≥30mm) and large zone diameters for oxacillin (≥13mm). These isolates are susceptible to both agents – this is the traditional wild-type phenotype for S. aureus. The lower left-hand quadrant represents isolates resistant to both drugs; these would be the MRSA isolates. The greatest number of isolates is in the upper left-hand section of the graph. Such isolates are resistant to penicillin (to the left of the red line), but susceptible to oxacillin (above the red line). This is the classical phenotype for beta-lactamase (penicillinase) producing S. aureus.
Fortunately, there are no isolates in the lower right-hand quadrant. Such isolates would have the phenotype penicillin-susceptible and oxacillin-resistant. Microbiologically, such results would be exceedingly rare, and most likely attributable to an error in the laboratory (poor quality reagents, test performance, measurement, transcription, etc.)
Thus a scatterplot can be useful both in the description of molecular mechanisms of resistance, as well as in the detection of unusual or impossible phenotypes suggestive of possible errors. Click on “Continue” to return to the main data analysis screen.
To obtain the scatterplot that you just reviewed, you will need quantitative data (zone diameters and/or MIC values). If you do not have these measurements, you can still do a this kind of comparison, but using the test interpretations rather than the measurements.
Click on “Analysis type”. In the middle of the screen, the current select is “Test measurements”. Click on “Test interpretations”. Click on “OK” and “Begin analysis”.
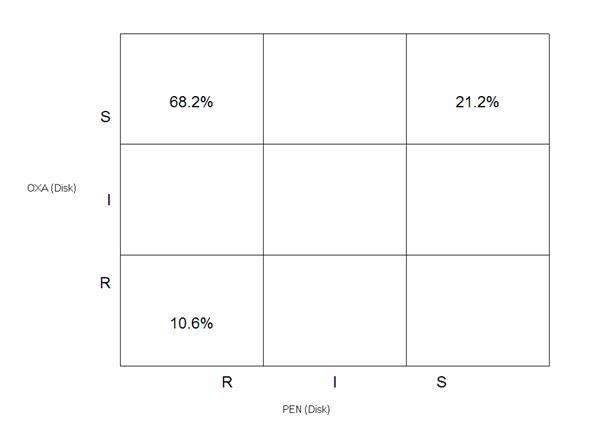
In this output, you see the same kind of results seen in the quantitative scatterplot, but using the R, I, S categories. You will again observe the three major subtypes of S. aureus in this chart. 21.2% of the isolates have the wild-type phenotype (PEN-S, OXA-S), 68.2% have the penicillinase-producer phenotype (PEN-R, OXA-S), and 10.6% have the MRSA phenotype (PEN-R, OXA-R). Click on “Continue” to return to the data analysis screen.
Click on “Analysis type”, and choose the following scatterplot options: X-Axis = Oxacillin-Disk and Y-Axis = Ciprofloxacin Disk. Do this scatterplot both by Test measurements and by Test interpretation to get the following two graphs.
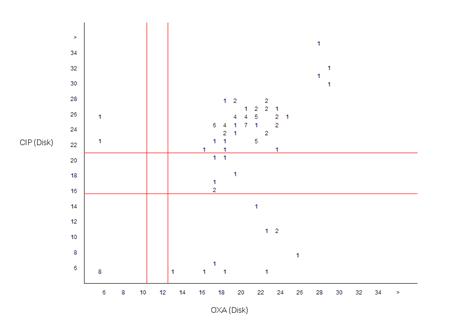
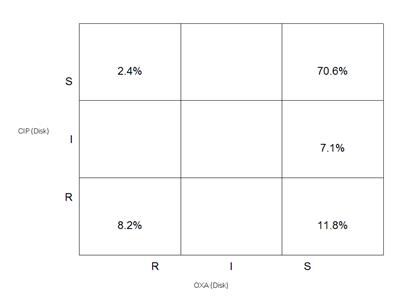
In the previous example, the two drugs selected (penicillin and oxacillin) were from the same class of drugs (beta-lactams). Such a comparison is useful for studying mechanisms of resistance and quality assurance (finding unlikely resistance phenotypes). In this example, the antibiotics oxacillin and ciprofloxacin are from two different classes of drugs with distinct mechanisms of resistance, so the graphs give information about cross-resistance and genetic linkages between classes. For example, in this case you see that most MRSA isolates (oxacillin-resistant) are also resistant to ciprofloxacin, a commonly observed epidemiological finding in many institutions.
Besides their value in studies of molecular epidemiology, studies of cross-resistance are useful to pharmacists and clinicians in developing policies for first-line and second-line treatment alternatives and to infection control staff in describing distinct clones causing hospital-acquired infections.
Part 6. Antimicrobial resistance profiles
The final analysis that covered in this tutorial is the study of multi-resistance patterns using the analysis “Resistance profiles”. Click on “Analysis type”, and select “Resistant profiles”. Because you are analyzing only one month of data, change the option “Month” in the middle of the screen to “Day”.
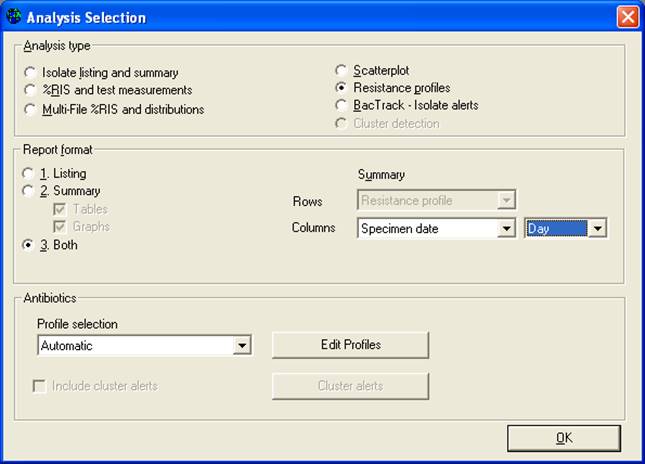
Then “OK” and “Begin analysis”. The following isolate listing shows the bottom portion of the output.
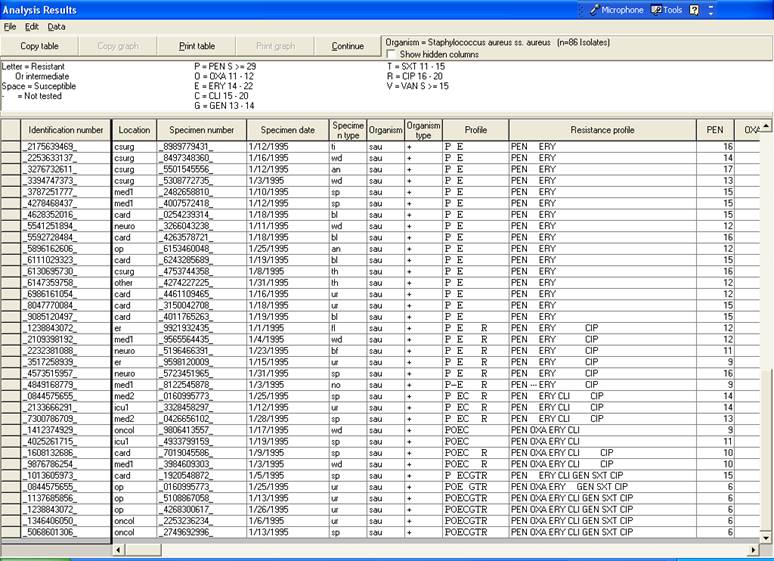
This listing is very similar to the one you did in Data analysis 1 with the analysis option “Isolate listing and summary”. But there are two additional columns: “Profile” (using one-letter antibiotic codes) and “Resistance profile” (using three-letter antibiotic codes). These columns indicate the multi-resistance pattern of the isolates. The profiles indicate the drugs to which the isolate is either resistant or intermediate (i.e. non-susceptible). For example “PE” = “PEN ERY” = isolates non-susceptible to penicillin and erythromycin, but susceptible to the other drugs requested.
Completely susceptible isolates appear at the top of the listing, followed by isolates non-susceptible to one drug, then to two drugs, etc. Multi-resistant strains appear at the bottom of the listing. This analysis thus categorizes all of the observed isolates according to resistance phenotype. If a patient has multiple isolates, you can see whether the isolates have the same resistance phenotype, or whether the phenotype changes over time, for example accumulating mutations, resistance genes, and plasmids. By examining the dates and the room numbers, you may also detect possible outbreaks of certain strains of S. aureus as defined by their multi-resistance pattern.
Click on “Continue” to see a summary of this list.
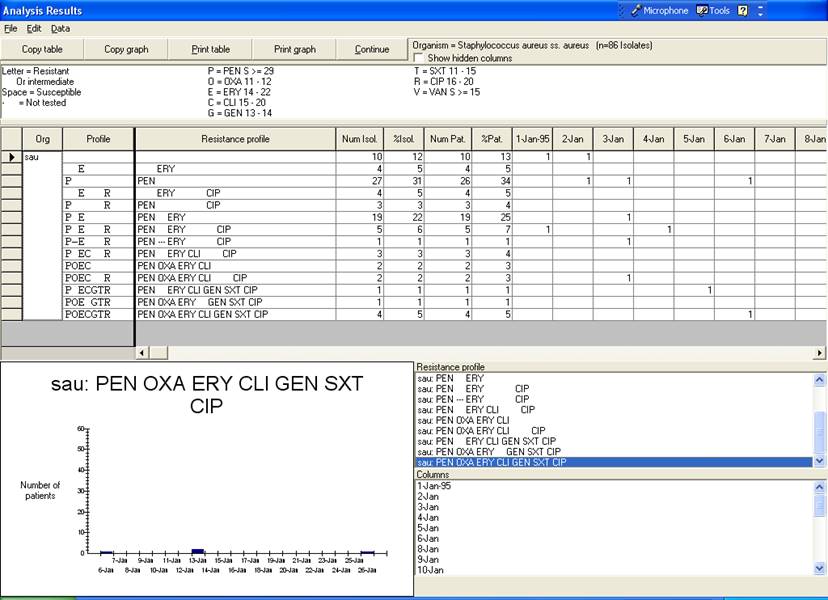
This output summarizes findings for all resistance phenotypes observed in the database. In this sample database, you will see that the most common phenotype is pan-susceptible (susceptible to all drugs requested) followed by “PEN” (non-susceptible to penicillin). Among MRSA isolates, the most common phenotype is “PEN OXA ERY CLI GEN SXT CIP”, in other words non-susceptible to all drugs requested with the exception of vancomycin. The day-by-day distribution of this MRSA phenotype is shown in the graph.
Click on “Continue” to return to the main menu.
In this analysis, WHONET did not analyze results for all of the drugs tested again S. aureus. Instead, WHONET used the drugs indicated in the laboratory configuration. To see and modify the drugs used for this analysis, click on “Analysis type” and “Edit profiles”.
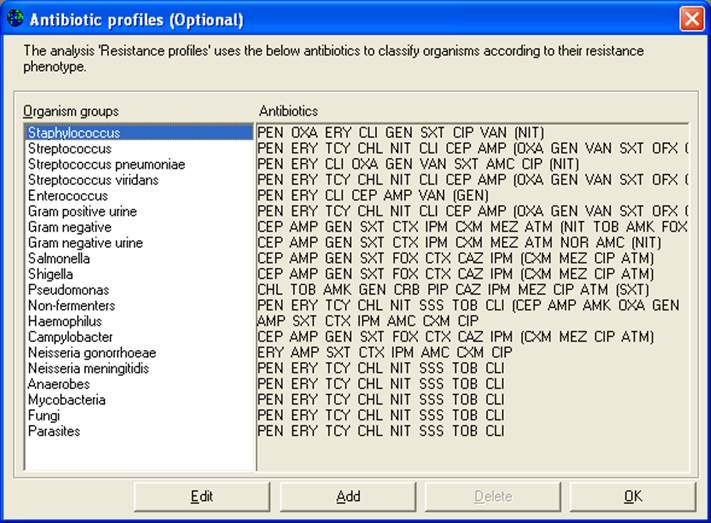
This list indicates which antimicrobials will be used, by default, for each organism group. For the item “Staphylococcus”, click on “Edit”.
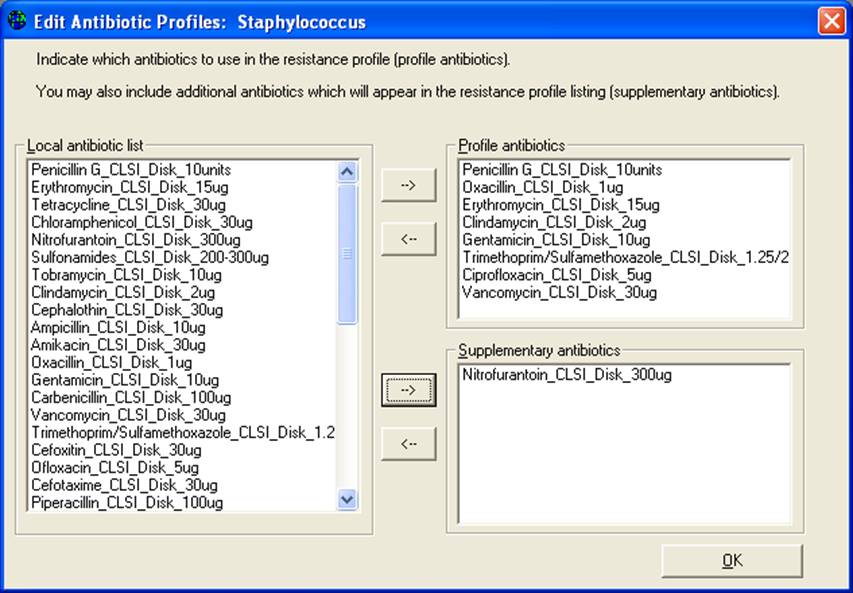
On this screen, you can select the drugs to use for the resistance profiles. Drugs in the box “Profile antibiotics” will be used to define the resistance phenotype. Drugs appearing under “Supplementary antibiotics” will appear in the line-listing, but are not used to define the resistance phenotype.
Note: Any modifications that you make to the profile antibiotics while you are in the Data analysis area of WHONET will be forgotten as soon as you leave WHONET. Any edits that you do here are temporary. To save the changes permanently for future use, you will need to use the WHONET configuration program. To do this, you will need to go back to the main WHONET screen and select “Modify laboratory” and “Antibiotics”. You will see a button called “Profiles”. After you change the profile antibiotics, click on “Save” to save them into your laboratory configuration.
Part 7. What next?
In the tutorials Data analysis 1 and Data analysis 2, you learned about the most commonly used WHONET analysis options and features. There are additional useful features covered in the following tutorials:
Expert System
Macros and Excel Reports
Cluster Detection with SaTScan (not activated yet, pending publication)